Domestikationsgenomik
Wildpflanzen wurden von den ersten Bauern domestiziert und verbreiteten sich mit der Landwirtschaft auf der ganzen Welt. Dabei hat sich ihr Erbgut deutlich verändert. Die Arbeitsgruppe Domestikationsgenomik erforscht die Prozesse Domestikation und Adaptation und ihrer Wechselwirkungen mit genetischer Diversität bei Kulturpflanzen und ihren verwandten Wildformen. Wir konzentrieren uns überwiegend auf die Getreide der gemäßigten Breiten: Gersten, Weizen, Roggen und Hafer. Die Bundeszentrale Ex-situ-Genbank mit tausenden Mustern von Wild- und Kulturformen ist für uns eine Ressource von unschätzbarem Wert. Unsere wichtigsten Forschungsziele sind:
- die Verwandtschaftsverhältnisse zwischen Kulturpflanzen und ihren wilden Ausgangsformen zu verstehen;
- die demografische Entwicklung und Anpassung der Getreide nachzuvollziehen, welche sich von Domestikationsursprung im Fruchtbaren Halbmond nach Europe ausgebreitet haben;
- die Auswirkungen der Domestikation auf molekularer Ebene zu entschlüsseln, z.B. auf die genetische Diversität sowie auf die Expression und Regulation von Genen.
Dazu wenden wir Methoden der Populationsgenetik und Genominformatik auf Sequenzdatensätze, genetische Markerdaten und Genexpressionsmatrizen an. Außerdem entwickeln wir genomische Sequenzressourcen und Kartierungsdaten für eine unvoreingenommene Bewertung der genetischen Diversität in hochdiversen Kulturpflanzensortimenten wie der IPK-Genbank. Wir tragen zu den IPK-Forschungsschwerpunkten “Genomdiversität und Evolution” und „Strategien zur Valorisierung pflanzengenetischer Ressourcen“ bei.
Dr. Martin Mascher ist Mitglied des Deutschen Zentrums für integrative Biodiversitätsforschung (iDiv) Halle-Jena-Leipzig.
scroll top
Projekte
Genomische Methoden zur Untersuchung der Evolution der Kulturpflanzen
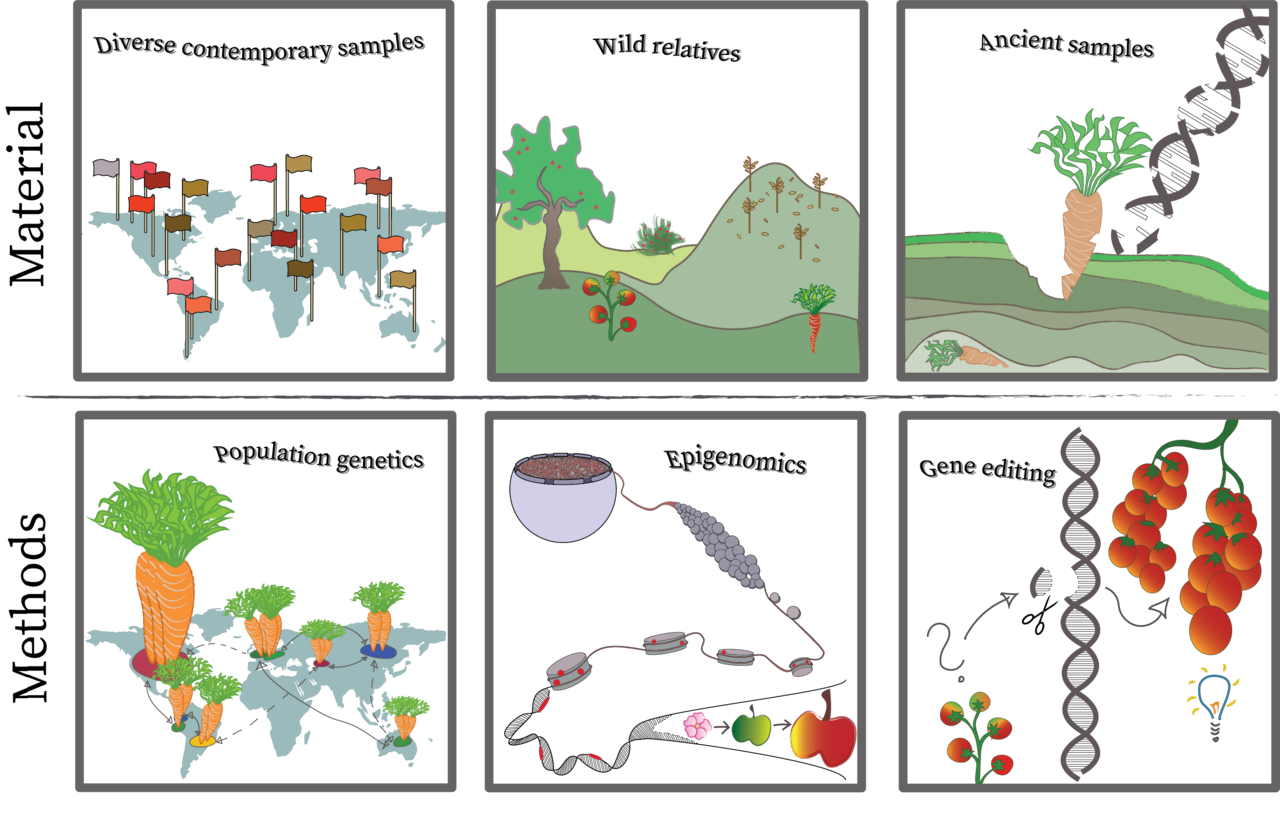
Wir wollen verstehen, wie unseren heutigen Kulturpflanzen aus ihren ursprünglichen Wildformen entstanden sind und sich auf der ganzen Welt ausgebreitet haben. Die Entwicklung fortgeschrittener genomischer Ressourcen und Werkzeuge ermöglicht es, genetische Kartierung und populationsgenetische Untersuchungen an diversen Kulturpflanzen und ihre wilden Verwandten durchführen, um die molekularen Grundlagen der Domestikation zu ergründen. Evolutionsgenetische Studien an Nutzpflanzen beruhen auf hochqualitativen Referenzsequenzen für Kultur- und Wilformen. Darauf aufbauend können Kulturpflanzensortimente (wie z.B. die IPK-Genbank) genomisch charakterisiert werden. Anschließend bietet das Methodenrepertoire der Archäogenetik, Epigenomik und Geneditierung vielfältige Möglichkeiten zu tiefergehender Forschung. Unsere Gruppe konzentriert sich auf genomische Methoden, aber wir arbeiten hinsichtlich anderer Aspekte mit institutsinternen, nationalen und internationalen Kooperationspartnern zusammen.
Literatur:
genomebiology.biomedcentral.com/articles/10.1186/s13059-018-1528-8
Pangenome der Getreide
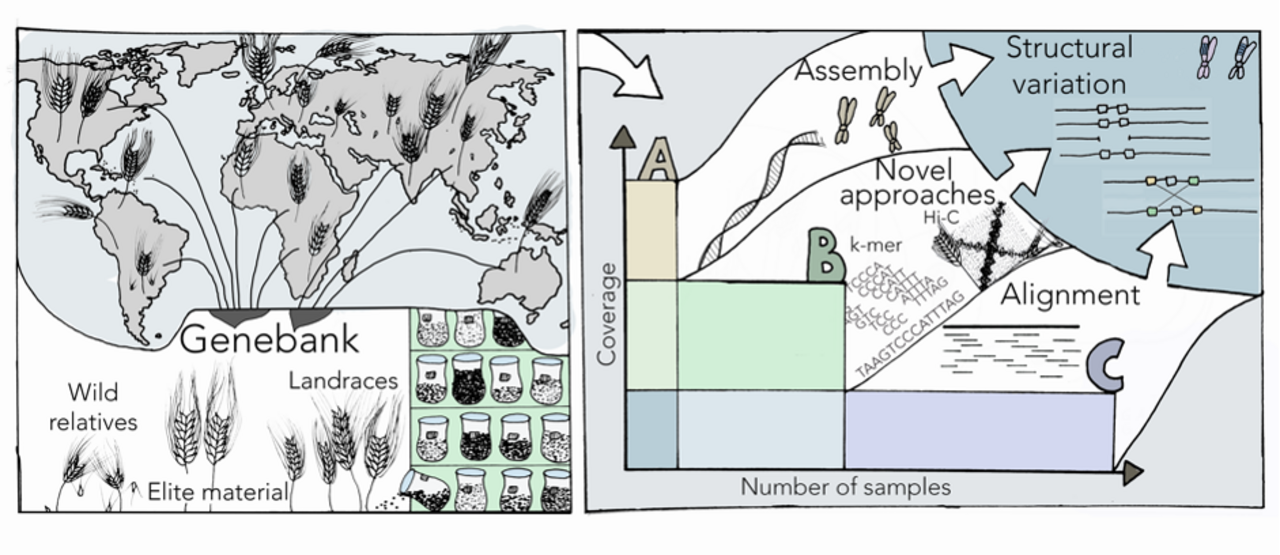
Das Forschungsbebiet Pangenomik befasst sich mit der vergleichenden Untersuchung von Genomsequenzen mehrerer Individuen einer Art. Pangenomik bei Nutzpflanzen konzentriert sich auf die Suche nach bisher unbekannter genetischer Variation, um die Züchtung neuer Sorten zu unterstützen. Genotypen für pangenomische Analysen werden aus großen Kulturpflanzensortimenten basierend auf genomweiten Markerdaten ausgewählt. Dies liefert in sich verschachtelte Kernsammlungen unterschiedlichen Umfangs als Repräsentanten der Diversität von Landrassen, Elitesorten und Wildformen. Referenzgenomsequenzen werden für wenige Schlüsselgenotypen (A) assembliert. Genomsequenzdaten mittlerer Dichte dienen zur referenzbasierten Suche nach struktureller Variation (B). Neuartige Methoden wie die Chromosomenstrukturanalyse und die Assoziationsgenetik mit kurzen Sequenzrepräsentanten werden bei der Suche nach chromosomalen Inversionen und der Verknüpfung von Genotyp und Phänotyp eingesetzt.
Literatur:
https://pubmed.ncbi.nlm.nih.gov/30446793/
https://academic.oup.com/dnaresearch/article/28/1/dsaa030/6117190
Archäogenetik and Genbankgenomik
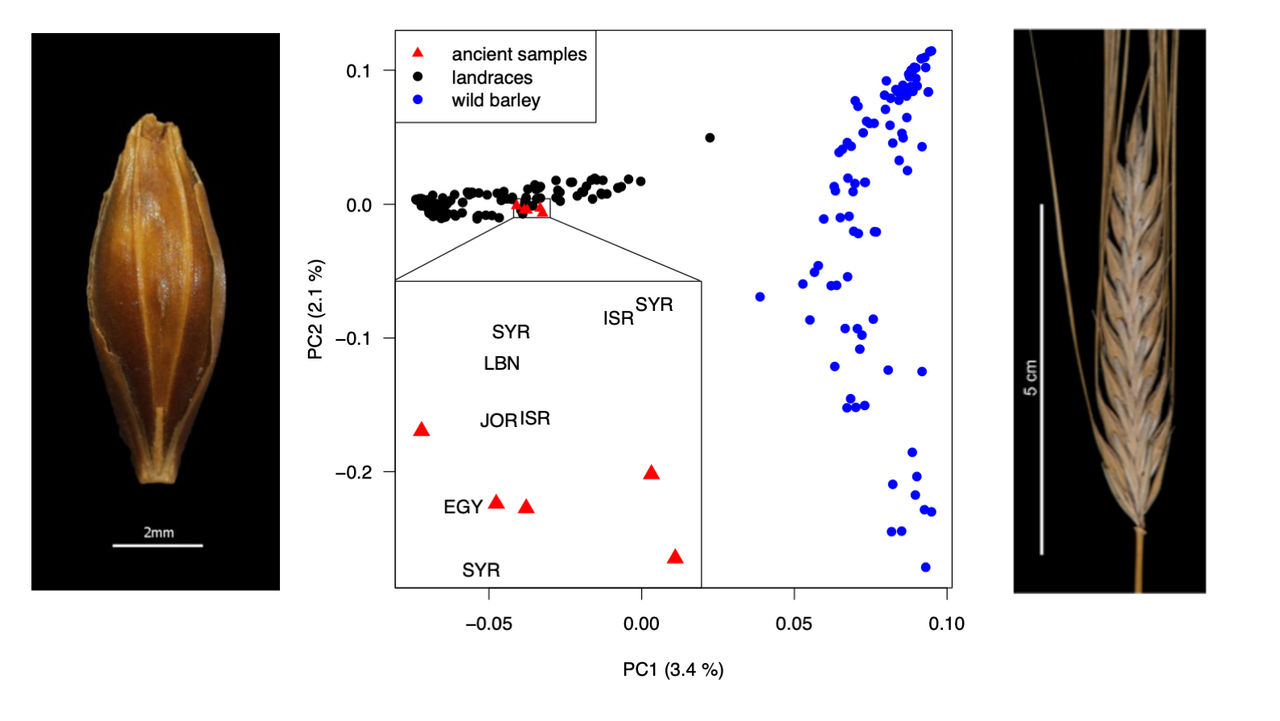
Das Wildgras Hordeum spontaneum wurde vor 10.000 Jahren im Fruchtbaren Halbmond domestiziert. Als Gerste wurde es eine der erste Nutzpflanzen der Menschheit. Wir haben Genomsequenzen von fünf 6000 Jahren alten Gerstenkörnern untersucht. Die Körner wurden von israelischen Archäologen in einer Höhle in der judäischen Wüste in der Nähe des Toten Meeres entdeckt. Ein Vergleich der alten Körner mit Sequenzdaten einer Gruppe diverser Gerstenmuster aus der IPK-Genbank zeigte, dass diese sehr nah mit Landrassen aus der südlichen Levante und Ägypten verwandt sind. Diese Ergebnisse deuten darauf hin, dass Gerstenlandrassen im heutigen Israel genetische Kontinuität über Jahrtausende zeigen. Wir fanden aber auch Hinweise auf Genfluss zwischen domestizierten Populationen und lokalen Wildformen. Die Abbildung verortet die fünf alten Proben in Diversitätsraum der modernen Gerste. Das Ährenmuster gehört zu einer der IPK-Genbankakzessionen, die am nächsten mit den alten Proben verwandt sind.
Literatur:
scroll top
Mitarbeitende
scroll top
Publikationen
Avni R, Kamal N, Bitz L, Jellen E N, Bekele W A, Angessa T T, Auvinen P, Bitz O, Boyle B, Canales F J, Carlson C H, Chapman B, Chawla H S, Chen Y, Copetti D, Correia de Lemos S, Dang V, Eichten S R, Klos K E, Fenn A M, Fiebig A, Fu Y-B, Gundlach H, Gupta R, Haberer G, He T, Herrmann M H, Himmelbach A, Howarth C J, Hu H, Isidro Y Sánchez J, Itaya A, Jannink J-L, Jia Y, Kaur R, Knauft M, Langdon T, Lux T, Marmon S, Marosi V, Mayer K F X, Michel S, Nandety R S, Nilsen K T, Paczos-Grzeda E, Pasha A, Prats E, Provart N J, Ravagnani A, Reid R W, Schlueter J A, Schulman A H, Sen T Z, Singh J, Singh M, Sirijovski N, Stein N, Studer B, Viitala S, Vronces S, Walkowiak S, Wang P, Waters A J, Wight C P, Yan W, Yao E, Zhang X-Q, Zhou G, Zhou Z, Tinker N A, Fiedler J D, Li C, Maughan P J, Spannagl M, Mascher M:
A pangenome and pantranscriptome of hexaploid oat. Nature 649 (2026) 131-139. https://dx.doi.org/10.1038/s41586-025-09676-7
Barzi S M A, Ahmadikhah A, Haghi R:
Comparative transcriptome analysis of two contrasting rice genotypes reveals the role of photosynthesis, CO2 fixation/metabolism and MAPK signaling pathways in response to drought stress. J. Plant Growth Reg. 45 (2026) 1537-1552. https://dx.doi.org/10.1007/s00344-025-11961-8
Chen Y, Copetti D, Kölliker R, Mascher M, Himmelbach A, Stein N, Studer B:
Chromosome-level haplotype-resolved assembly of highly heterozygous grass genomes with PhaseGrass. Nat. Commun. 17 (2026) 12. https://dx.doi.org/10.1038/s41467-025-66377-5
Devabhakthini N, Eichler-Löbermann B, Haghi R, Willner E, Dehmer K J, Kavka M:
Effect of phosphorus deficiency on biomass and root system architecture in diverse Medicago accessions. Front. Plant Sci. 17 (2026) 1812278. https://dx.doi.org/10.3389/fpls.2026.1812278
Ganji E, Harpke D, Himmelbach A, Lohwasser U:
Uncovering the phylogenetic relationships of lentil (Lens culinaris Medik.) accessions from Germany. Genet. Resour. Crop Evol. 73 (2026) 183. https://dx.doi.org/10.1007/s10722-026-02810-y
Haraldsson E B, Anokye M, Rütjes T, Toegelová H, Tulpová Z, Šimková H, Feng J-W, Mascher M, von Korff M:
A chromosome-scale genome assembly of Hordeum erectifolium: genomic, transcriptomic and anatomical adaptations to drought in a wild barley relative. New Phytol. 250 (2026) 2652-2669. https://dx.doi.org/10.1111/nph.71091
Jørgensen M E, Vequaud D, Wang Y, Andersen C B, Bayer M, Box A, Braune K B, Cai Y, Chen F, Cuesta-Seijo J A, Dong H, Fincher G B, Gojkovic Z, Huang Z, Jaegle B, Kale S M, Krsticevic F, Le Roux P-M, Lozier A, Lu Q, Mascher M, Murozuka E, Nakamura S, Simmelsgaard M U, Pedas P R, Pin P A, Podzimska-Sroka D, Sato K, Spannagl M, Rasmussen M W, Russell J, Schreiber M, Thomsen H C, Thomsen N W, Tulloch S, Voss C, Skadhauge B, Stein N, Willerslev E, Waugh R, Dockter C:
Postdomestication selection of MKK3 shaped seed dormancy and end-use traits in barley. www.science.org/stoken/author-tokens/ST-3031/full. Science 391 (2026) 90-95. https://dx.doi.org/10.1126/science.adx2022
Kale S M, Gundlach H, Gericke O, Kamal N, Almeida A, Pitra N, Price N, Haberer G, Lux T, Krsticevic F, Kemp O, de Bang L, Himmelbach A, Padmarasu S, Rabanus-Wallace M-T, Horáková L, Bačovský V, Nielsen K, Bjarnholt N, Micic N, Kruse-Andersen I, Møller B L, Janfelt C, Skadhauge B, Matthews P D, Mayer K F X, Stein N, Mascher M, Spannagl M, Feiner A, Braumann I:
Extensive variation between chromosomes of North American and European hop. Nat. Commun. 17 (2026) 4110. https://dx.doi.org/10.1038/s41467-026-72379-8
Karami-Moalem S, Ahmadikhah A, Nemati Z, Haghi R:
EMS-induced genomic variation and QTL-seq analysis of safflower spininess through whole genome sequencing (WGS). BMC Genomics 27 (2026) 115. https://dx.doi.org/10.1186/s12864-025-12488-8
Khazaei Z, Bagheri A, Lyskov D, Harpke D, Blattner F R:
Root stomata in Conium maculatum (Apiaceae): anatomically verified occurrence and a comparative survey across Apioideae. AoB Plants 18 (2026) plag001. https://dx.doi.org/10.1093/aobpla/plag001
Lev-Mirom Y, Ashkenazy N, Klymiuk V, Ens J, Wiebe K, Schuenemann V J, Eilam T, Kulikovsky S, Lavi B A, Günther T, Krugman T, Sela H, Klein E, Ganor A, Davidovich U, Porat R, Ben-Yosef E, David M, Frankin S, Ben-David R, Cavalet-Giorsa E, Krattinger S G, Korol A, Mascher M, Himmelbach A, Stein N, Krause J, Distelfeld A, Weiss E, Fahima T, Pozniak C J, Hübner S:
Ancient grains illuminate the mosaic origin of domesticated wheat. Nat. Plants 12 (2026) 954-963. https://dx.doi.org/10.1038/s41477-026-02283-y
Sabzeali F, Ahmadikhah A, Farrokhi N, Haghi R:
Transcriptome analysis of safflower (Carthamus tinctorius L.) reveals the roles of osmotic adjustment and regulatory mechanisms in response to drought stress. BMC Plant Biol. 26 (2026) 503. https://dx.doi.org/10.1186/s12870-026-08366-4
Sardouei-Nasab S, Mahdavinasab F, Haghi R, Mohammadi-Nejad G, Nikbakht-Dehkordi A, Nemati Z:
Integrative GWAS and transcriptomic analyses identify candidate genes associated with oil content and fatty acid composition in safflower. BMC Plant Biol. 26 (2026) 372. https://dx.doi.org/10.1186/s12870-025-08024-1
Spanner R, Sallam A H, Guo Y, Jayakodi M, Himmelbach A, Fiebig A, Simmons J, Bethke G, Lee Y, Pacheco Arge L W, Qiu Y, Badea A, Baum M, Belzile F, Ben-David R, Brueggeman R, Case A, Cattivelli L, Davis M, Dockter C, Doležel J, Dreiseitl A, Gavin R, Glick L, Greiner S, Hamilton R, Hayes P M, Heisel S, Henson C, Kilian B, Komatsuda T, Li C, Liu C, Mahalingam R, Maruschewski M, Matny O, Maurer A, Mayer K F X, Mayrose I, Moscou M, Muehlbauer G J, Oono Y, Ordon F, Özkan H, Pecinka A, Perovic D, Pillen K, Pourkheirandish M, Russell J, Šafář J, Salvi S, Sanchez-Garcia M, Sato K, Schmutzer T, Scholz U, Scott J, Singh Brar G, Smith K P, Sorrells M E, Spannagl M, Stein N, Tondelli A, Tuberosa R, Tucker J, Turkington T, Valkoun J, Verma R P S, Vinje M A, von Korff M, Walling J G, Waugh R, Wise R P, Wulff B B H, Yang S, Zhang G, Morrell P L, Mascher M, Steffenson B J:
Whole-genome resequencing of the wild barley diversity collection: a resource for identifying and exploiting genetic variation for cultivated barley improvement. G3 Genes Genom. Genet. 16 (2026) jkaf261. https://dx.doi.org/10.1093/g3journal/jkaf261
Tao X-Y, Tan C, Liu Y, Wang Y, Raza A, He J, Wang L, Xia K, Yan Y, Liao S, Jiang W, Qu Y, Xu B, Zhou Y, Yang X, Roy S, Denton M, Tucker M, Able J, Gilliham M, Langridge P, Hu H-W, He J-Z, Zhao C, Zhou M, Mir R R, Chitikineni A, Li C, Zhang P, Trethowan R, Ding Y, Zhang J, Franks P, Wu Q, Ye L, Wang Y, Wu F, Zhang G, Cai S, Shen Q, Jiang H, Chen G, Zhou Y, Song C, Zhang Y, Ma W, Li X, Guo W, Zeng J, He X, Liu W, Feng X, Zhang R, Li G, Cao X, Liu S, Liu Q, Chen J, Zhang X, Liu L, Zeng F, Deng F, Qin Y, Zhu X, Shabala S, Tee E E, Schnurbusch T, Mascher M, Wu H, Spannagl M, Xin M, Gou J, Brar G, Xue D, Wang W, Reynolds M, Zhao Y, Liu Z, Krugman T, Sakata Y, Yee S, Chan K, Pogson B, Zhang Y, Marchant D B, Siddique K H M, Fang X, Chen A, Henry R, Boden S A, Varshney R K, Xu S-C, Xu X, Chen Z-H:
The potential of wheat spatial omics. Nat. Genet. 58 (2026) 962-973. https://dx.doi.org/10.1038/s41588-026-02542-w
Wu W, Jiang C, Gao G, Mitani-Ueno N, Guo Y, Kan J, Xiong Y, Mascher M, Ni Z, Ma J F, Wang K, Yang P:
Spatial regulation of silicon accumulation in peduncle confers sheathed spike in barley. Plant Biotechnol. J. (2026) Epub ahead of print. https://dx.doi.org/10.1111/pbi.70666
Zhang H, Windhorst A, Bornhofen E, Tulpová Z, Novak P, Macas J, Šimková H, Nadzieja M, Kim J M, Cram D, Cao Y, Konkin D J F, Sass O, Welna G, Himmelbach A, Mascher M, Link W, Kwon S-J, Yang T-J, Andersen S U, Jayakodi M:
Allelic variation at a single locus distinguishes spring and winter faba beans. Nat. Genet. 58 (2026) 655-663. https://dx.doi.org/10.1038/s41588-026-02524-y
Bekele W A, Avni R, Birkett C L, Itaya A, Wight C P, Bellavance J, Brodführer S, Canales F J, Carlson C H, Fiebig A, Li Y, Michel S, Nandety R S, Waring D J, Arbelaez J D, Beattie A D, Caffe M, del Blanco I A, Fiedler J D, Gupta R, Gutierrez L, Harris J C, Harrison S A, Herrmann M H, Huang Y-F, Isidro Y Sanchez J, McMullen M S, Mitchell Fetch J W, Nilsen K T, Parkin I A P, Peng Y, Smith K P, Sutton T, Yan W, Zwer P, Diederichsen A, Esvelt Klos K, Fu Y-B, Howarth C J, Jannink J-L, Jellen E N, Langdon T, Maughan P J, Paczos-Grzeda E, Prats E, Sen T Z, Mascher M, Tinker N A:
Global genomic population structure of wild and cultivated oat reveals signatures of chromosome rearrangements. Nat. Commun. 16 (2025) 9486. https://dx.doi.org/10.1038/s41467-025-57895-3
Câmara A S, Mascher M:
Understanding and simulating the dynamics of a polymer-like chromatin. In: Baroux C, Tatout C (Eds.): Methods for plant nucleus and chromatin studies: methods and protocols. (Series: Methods in molecular biology, Vol. 2873) New York: Humana Press (2025) 978-1-0716-4228-3, 283-302. https://dx.doi.org/10.1007/978-1-0716-4228-3_16
Cho W, Feng J, Knauft M, Albrecht S, Himmelbach A, Otto L-G, Mascher M:
An annotated haplotype-resolved genome sequence assembly of diploid German chamomile, Matricaria chamomilla. Sci. Data 12 (2025) 358. https://dx.doi.org/10.1038/s41597-025-04688-4
Feng J-W, Pidon H, Cuacos M, Lux T, Himmelbach A, Haghi R, Fuchs J, Haberer G, Kuo Y-T, Guo Y, Jayakodi M, Toegelová H, Harpke D, Knauft M, Fiebig A, Maruschewski M, Ronen M, Sharon A, Šimková H, Mayer K F X, Spannagl M, Kumlehn J, Heckmann S, Houben A, Blattner F R, Stein N, Mascher M:
A haplotype-resolved pangenome of the barley wild relative Hordeum bulbosum. Nature 645 (2025) 429-438. https://dx.doi.org/10.1038/s41586-025-09270-x
Guo W, Schreiber M, Marosi V B, Bagnaresi P, Jørgensen M E, Braune K B, Chalmers K, Chapman B, Dang V, Dockter C, Fiebig A, Fincher G B, Fricano A, Fuller J, Haaning A, Haberer G, Himmelbach A, Jayakodi M, Jia Y, Kamal N, Langridge P, Li C, Lu Q, Lux T, Mascher M, Mayer K F X, McCallum N, Milne L, Muehlbauer G J, Nielsen M T S, Padmarasu S, Pedas P R, Pillen K, Pozniak C, Rasmussen M W, Sato K, Schmutzer T, Scholz U, Schüler D, Šimková H, Skadhauge B, Stein N, Thomsen N W, Voss C, Wang P, Wonneberger R, Zhang X-Q, Zhang G, Cattivelli L, Spannagl M, Bayer M, Simpson C, Zhang R, Waugh R:
A barley pan-transcriptome reveals layers of genotype-dependent transcriptional complexity. Nat. Genet. 57 (2025) 441-450. https://dx.doi.org/10.1038/s41588-024-02069-y
Guo Y:
A haplotype-based evolutionary history of barley domestication. (PhD Thesis) Halle/S., Martin-Luther-Universität Halle-Wittenberg, Naturwissenschaftliche Fakultät III Agrar- und Ernährungswissenschaften, Geowissenschaften und Informatik (2025) 122 pp.
Guo Y, Jayakodi M, Himmelbach A, Ben-Yosef E, Davidovich U, David M, Hartmann-Shenkman A, Kislev M, Fahima T, Schuenemann V J, Reiter E, Krause J, Steffenson B J, Stein N, Weiss E, Mascher M:
A haplotype-based evolutionary history of barley domestication. Nature 647 (2025) 680-688. https://dx.doi.org/10.1038/s41586-025-09533-7
Heilmann P G, Grosch E, Frisch M, Herrmann M, Beuch S, Kurra V, Mascher M, Avni R, Oldach K, Röhrs I, Hanemann A, Mehta R R, Reinbrecht C, Serfling A, Stahl A, Stucke M, Abbadi A, Kox T, Zenke-Philippi C:
Haplotype-based autoencoders can reduce the dataset dimension and estimate haplotype block effects in different crop species. BMC Bioinformatics 26 (2025) 289. https://dx.doi.org/10.1186/s12859-025-06323-w
Jayakodi M, Shim H, Mascher M:
What are we learning from plant pangenomes? Annu. Rev. Plant Biol. 76 (2025) 663-686. https://dx.doi.org/10.1146/annurev-arplant-090823-015358
Khosravi S, Hinrichs R, Rönspies M, Haghi R, Puchta H, Houben A:
Epigenetic state and gene expression remain stable after CRISPR/Cas-mediated chromosomal inversions. New Phytol. 245 (2025) 2527-2539. https://doi.org/10.1111/nph.20403
Sabili Z, Rashidi-Monfard S, Haghi R, Kahrizi D:
Comparative analysis of simple sequence repeats and synteny across ten Oryza species: Implications for stress response and genetic diversity. Comput. Biol. Chem. 116 (2025) 108379. https://dx.doi.org/10.1016/j.compbiolchem.2025.108379
Sabzeali F, Ahmadikhah A, Farrokhi N, Haghi R:
Genome-wide identification and transcription profiling of safflower (Carthamus tinctorius L.) HD-ZIP gene family under water deficit. BMC Genomics 26 (2025) 874. https://dx.doi.org/10.1186/s12864-025-12060-4
Sharma R, Shaaf S, Neumann K, Civan P, Guo Y, Mascher M, David M, Al-Yassin A, Özkan H, Blake T, Hübner S, Castañeda-Álvarez N P, Grando S, Ceccarelli S, Baum M, Graner A, Coupland G, Pillen K, Weiss E, Mackay I J, Powell W, Kilian B:
On the origin of the late-flowering ppd-H1 allele in barley. Theor. Appl. Genet. 138 (2025) 246. https://dx.doi.org/10.1007/s00122-025-04981-1
Thudi M, Mascher M, Jayakodi M:
Pangenome charts the genomic path for wheat improvement. Trends Plant Sci. 30 (2025) 687-689. https://dx.doi.org/10.1016/j.tplants.2025.03.002
Vaghari S, Fazeli A, Haghi R, Shohani F:
Physiological and biochemical responses of Carthamus tinctorius L. under salt stress. Kuwait J. Sci. 52 (2025) 100464. https://doi.org/10.1016/j.kjs.2025.100464
White B, Lux T, Rusholme-Pilcher R, Juhász A, Kaithakottil G, Duncan S, Simmonds J, Rees H, Wright J, Colmer J, Ward S, Joynson R, Coombes B, Irish N, Henderson S, Barker T, Chapman H, Catchpole L, Gharbi K, Bose U, Okada M, Handa H, Nasuda S, Shimizu K K, Gundlach H, Lang D, Naamati G, Legg E J, Bharti A K, Colgrave M L, Haerty W, Uauy C, Swarbreck D, Borrill P, Poland J A, Krattinger S G, Stein N, Mayer K F X, Pozniak C, Walkowiak S, Klymiuk V, Byrns B, Nilsen K, Ens J, Wiebe K, N’Diaye A, Hucl P J, Pozniak C J, Fu B X, Gao L, Delorean E, Koo D-H, Fritz A K, Poland J, Monat C, Himmelbach A, Fiebig A, Padmarasu S, Scholz U, Mascher M, Haberer G, Kassa M T, Fobert P, Kagale S, Brinton J, Ramirez-Gonzalez R H, Bevan M, McKenzie N, Steuernagel B, Kolodziej M C, Krattinger S G, Keller B, Wicker T, Thambugala D, McCartney C A, Bandi V, Siri J N, Gutwin C, Aquino C, Hatakeyama M, Copetti D, Halstead-Nussloch G, Paape T, Shimizu-Inatsugi R, Shimizu K K, Ban T, Kawaura K, Tameshige T, Tsuji H, Venturini L, Clark M, Clavijo B, Fosker C, Accinelli G G, Heavens D, Krasileva K, Gardner K A, Fradgley N, Percival-Alwyn L, Cockram J, Gutierrez-Gonzalez J, Muehlbauer G, Koh C S, Sharpe A G, Deek J, Costamagna A C, Kanamori H, Kobayashi F, Tanaka T, Wu J, Handa H, Kuo T, Sese J, Murata K, Nabeka Y, Nasuda S, Juliana P, Singh R, Budak H, Small I, Melonek J, Cloutier S, Keeble-Gagnère G, Tibbets J, Legg E, Bharti A, Langridge P, Chalmers K, Distelfeld A, Spannagl M, Hall A, Wheat Genome Project:
De novo annotation reveals transcriptomic complexity across the hexaploid wheat pan-genome. Nat. Commun. 16 (2025) 8538. https://dx.doi.org/10.1038/s41467-025-64046-1
Yuan Z, Rembe M, Mascher M, Stein N, Himmelbach A, Jayakodi M, Börner A, Oldach K, Jahoor A, Jensen J D, Rudloff J, Dohrendorf V-E, Kuhfus L P, Dyrszka E, Conte M, Hinz F, Trouchaud S, Reif J C, El Hanafi S:
High-quality phenotypic and genotypic dataset of barley genebank core collection to unlock untapped genetic diversity. GigaScience 14 (2025) giae121. https://dx.doi.org/10.1093/gigascience/giae121
Zhang H:
Improved genome and population genomics unveil the winter hardiness and breeding signatures in faba bean. (PhD Thesis, kumulativ) Göttingen, Georg-August-Universität Göttingen, Fakultät für Agrarwissenschaften (2025) 145 pp.
Câmara A S, Kubalová I, Schubert V:
Helical chromonema coiling is conserved in eukaryotes. Plant J. 118 (2024) 1284-1300. https://dx.doi.org/10.1111/tpj.16484
Cavalet-Giorsa E, González-Muñoz A, Athiyannan N, Holden S, Salhi A, Gardener C, Quiroz-Chávez J, Rustamova S M, Elkot A F, Patpour M, Rasheed A, Mao L, Lagudah E S, Periyannan S K, Sharon A, Himmelbach A, Reif J C, Knauft M, Mascher M, Stein N, Chayut N, Ghosh S, Perovic D, Putra A, Perera A B, Hu C-Y, Yu G, Ahmed H I, Laquai K D, Rivera L F, Chen R, Wang Y, Gao X, Liu S, Raupp W J, Olson E L, Lee J-Y, Chhuneja P, Kaur S, Zhang P, Park R F, Ding Y, Liu D-C, Li W, Nasyrova F Y, Dvorak J, Abbasi M, Li M, Kumar N, Meyer W B, Boshoff W H P, Steffenson B J, Matny O, Sharma P K, Tiwari V K, Grewal S, Pozniak C J, Chawla H S, Ens J, Dunning L T, Kolmer J A, Lazo G R, Xu S S, Gu Y Q, Xu X, Uauy C, Abrouk M, Bougouffa S, Brar G S, Wulff B B H, Krattinger S G:
Origin and evolution of the bread wheat D genome. Nature 633 (2024) 848-855. https://dx.doi.org/10.1038/s41586-024-07808-z
Chen Y, Kölliker R, Mascher M, Copetti D, Himmelbach A, Stein N, Studer B:
An improved chromosome-level genome assembly of perennial ryegrass (Lolium perenne L.). GigaByte 2024 (2024) gigabyte112. https://dx.doi.org/10.46471/gigabyte.112
Cho W, Jang W, Shim H, Kim J, Oh Y, Park J Y, Kim Y C, Lee J-W, Jo I-H, Lee M, Gil J, Mascher M, Jayakodi M, Liao X, Xu J, Dou D, Lee Y, Yang T-J:
High-resolution genetic map and SNP chip for molecular breeding in Panax ginseng, a tetraploid medicinal plant. Hortic. Res. 11 (2024) uhae257. https://dx.doi.org/10.1093/hr/uhae257
Fusi R, Milner S G, Rosignoli S, Bovina R, De Jesus Vieira Teixeira C, Lou H, Atkinson B S, Borkar A N, York L M, Jones D H, Sturrock C J, Stein N, Mascher M, Tuberosa R, O'Connor D, Bennett M J, Bishopp A, Salvi S, Bhosale R:
The auxin efflux carrier PIN1a regulates vascular patterning in cereal roots. New Phytol. 244 (2024) 104-115. https://dx.doi.org/10.1111/nph.19777
Gao G, Yan L, Cai Y, Guo Y, Jiang C, He Q, Tasnim S, Feng Z, Liu J, Zhang J, Komatsuda T, Mascher M, Yang P:
Most Tibetan weedy barleys originated via recombination between Btr1 and Btr2 in domesticated barley. Plant Commun. 5 (2024) 100828. https://dx.doi.org/10.1016/j.xplc.2024.100828
Garg V, Bohra A, Mascher M, Spannagl M, Xu X, Bevan M W, Bennetzen J L, Varshney R K:
Unlocking plant genetics with telomere-to-telomere genome assemblies. Nat. Genet. 56 (2024) 1788-1799. https://dx.doi.org/10.1038/s41588-024-01830-7
Güngör E, Savary J, Adema K, Dijkhuizen L W, Keilwagen J, Himmelbach A, Mascher M, Koppers N, Bräutigam A, Van Hove C, Riant O, Nierzwicki-Bauer S, Schluepmann H:
The crane fly glycosylated triketide δ-lactone cornicinine elicits akinete differentiation of the cyanobiont in aquatic Azolla fern symbioses. Plant Cell Environ. 47 (2024) 2675-2692. https://dx.doi.org/10.1111/pce.14907
Huang Y, Maurer A, Giehl R F H, Zhao S, Golan G, Thirulogachandar V, Li G, Zhao Y, Trautewig C, Himmelbach A, Börner A, Jayakodi M, Stein N, Mascher M, Pillen K, Schnurbusch T:
Dynamic phytomeric growth contributes to local adaptation in barley. Mol. Biol. Evol. 41 (2024) msae011. https://dx.doi.org/10.1093/molbev/msae011
Jayakodi M, Lu Q, Pidon H, Rabanus-Wallace M T, Bayer M, Lux T, Guo Y, Jaegle B, Badea A, Bekele W, Brar G S, Braune K, Bunk B, Chalmers K J, Chapman B, Jørgensen M E, Feng J-W, Feser M, Fiebig A, Gundlach H, Guo W, Haberer G, Hansson M, Himmelbach A, Hoffie I, Hoffie R E, Hu H, Isobe S, König P, Kale S M, Kamal N, Keeble-Gagnère G, Keller B, Knauft M, Koppolu R, Krattinger S G, Kumlehn J, Langridge P, Li C, Marone M P, Maurer A, Mayer K F X, Melzer M, Muehlbauer G J, Murozuka E, Padmarasu S, Perovic D, Pillen K, Pin P A, Pozniak C J, Ramsay L, Pedas P R, Rutten T, Sakuma S, Sato K, Schüler D, Schmutzer T, Scholz U, Schreiber M, Shirasawa K, Simpson C, Skadhauge B, Spannagl M, Steffenson B J, Thomsen H C, Tibbits J F, Nielsen M T S, Trautewig C, Vequaud D, Voss C, Wang P, Waugh R, Westcott S, Rasmussen M W, Zhang R, Zhang X-Q, Wicker T, Dockter C, Mascher M, Stein N:
Structural variation in the pangenome of wild and domesticated barley. Nature 636 (2024) 654-662. https://dx.doi.org/10.1038/s41586-024-08187-1
Jiang G, Koppolu R, Rutten T, Hensel G, Lundqvist U, Tandron Moya Y A, Huang Y, Rajaraman J, Poursarebani N, von Wirén N, Kumlehn J, Mascher M, Schnurbusch T:
Non-cell-autonomous signaling associated with barley ALOG1 specifies spikelet meristem determinacy. Curr. Biol. 34 (2024) 2344-2358. https://doi.org/10.1016/j.cub.2024.04.083
Kudapa H, Ghatak A, Barmukh R, Chaturvedi P, Khan A, Kale S, Fragner L, Chitikineni A, Weckwerth W, Varshney R K:
Integrated multi-omics analysis reveals drought stress response mechanism in chickpea (Cicer arietinum L.). Plant Genome 17 (2024) e20337. https://dx.doi.org/10.1002/tpg2.20337
Mascher M, Jayakodi M, Shim H, Stein N:
Promises and challenges of crop translational genomics. Nature 636 (2024) 585-593. https://dx.doi.org/10.1038/s41586-024-07713-5
Mascher M, Marone M P, Schreiber M, Stein N:
Are cereal grasses a single genetic system? rdcu.be/dErP0. Nat. Plants 10 (2024) 719-731. https://dx.doi.org/10.1038/s41477-024-01674-3
Moghadam A, Taghizadeh M S, Haghi R, Tahmasebi A, Niazi A, Ebrahimie E:
Exploring novel insights: Methyl jasmonate treatment reveals novel lncRNA-mediated regulation of secondary metabolite biosynthesis pathways in Echinacea purpurea. Food Biosci. 57 (2024) 103457. https://doi.org/10.1016/j.fbio.2023.103457
Schreiber M, Jayakodi M, Stein N, Mascher M:
Plant pangenomes for crop improvement, biodiversity and evolution. Nat. Rev. Genet. 25 (2024) 563-577. https://dx.doi.org/10.1038/s41576-024-00691-4
Sharma D, Avni R, Gutierrez-Gonzalez J, Kumar R, Sela H, Prusty M R, Shatil-Cohen A, Molnár I, Holušová K, Said M, Doležel J, Millet E, Khazan-Kost S, Landau U, Bethke G, Sharon O, Ezrati S, Ronen M, Maatuk O, Eilam T, Manisterski J, Ben-Yehuda P, Anikster Y, Matny O, Steffenson B J, Mascher M, Brabham H J, Moscou M J, Liang Y, Yu G, Wulff B B H, Muehlbauer G, Minz-Dub A, Sharon A:
A single NLR gene confers resistance to leaf and stripe rust in wheat. Nat. Commun. 15 (2024) 9925. https://dx.doi.org/10.1038/s41467-024-54068-6
Šimková H, Câmara A S, Mascher M:
Hi-C techniques: from genome assemblies to transcription regulation. J. Exp. Bot. 75 (2024) 5357-5365. https://dx.doi.org/10.1093/jxb/erae085
Thirulogachandar V, Govind G, Hensel G, Kale S M, Kuhlmann M, Eschen-Lippold L, Rutten T, Koppolu R, Rajaraman J, Palakolanu S R, Seiler C, Sakuma S, Jayakodi M, Lee J, Kumlehn J, Komatsuda T, Schnurbusch T, Sreenivasulu N:
HOMEOBOX2, the paralog of SIX-ROWED SPIKE1/HOMEOBOX1, is dispensable for barley spikelet development. J. Exp. Bot. 75 (2024) 2900-2916. https://dx.doi.org/10.1093/jxb/erae044
Tikhenko N, Haupt M, Fuchs J, Perovic D, Himmelbach A, Mascher M, Houben A, Rutten T, Nagel M, Tsvetkova N V, Sehmisch S, Börner A:
Major chromosome rearrangements in intergeneric wheat × rye hybrids in compatible and incompatible crosses detected by GBS read coverage analysis. Sci. Rep. 14 (2024) 11010. https://dx.doi.org/10.1038/s41598-024-61622-1
Yuan Z, Rembe M, Mascher M, Stein N, Jayakodi M, Börner A, Oldach K, Jahoor A, Jensen J D, Rudloff J, Dohrendorf V-E, Kuhfus L P, Dyrszka E, Conte M, Hinz F, Trouchaud S, Reif J C, El Hanafi S:
Capitalizing genebank core collections for rare and novel disease resistance loci to enhance barley resilience. J. Exp. Bot. 75 (2024) 5940-5954. https://dx.doi.org/10.1093/jxb/erae283
Ahmadli U, Kalidass M, Crhak Khaitova L, Fuchs J, Cuacos M, Demidov D, Zuo S, Pecinkova J, Mascher M, Ingouff M, Heckmann S, Houben A, Riha K, Lermontova I:
High temperature increases centromere-mediated genome elimination frequency and enhances haploid induction in Arabidopsis. Plant Commun. 4 (2023) 100507. https://dx.doi.org/10.1016/j.xplc.2022.100507
Brodführer S, Mohler V, Stadlmeier M, Okoń S, Beuch S, Mascher M, Tinker N A, Bekele W A, Hackauf B, Herrmann M H:
Genetic mapping of the powdery mildew resistance gene Pm7 on oat chromosome 5D. Theor. Appl. Genet. 136 (2023) 53. https://dx.doi.org/10.1007/s00122-023-04288-z
Câmara A S, Mascher M:
Consistencies and contradictions in different polymer models of chromatin architecture. Comput. Struct. Biotechnol. J. 21 (2023) 1084-1091. https://dx.doi.org/10.1016/j.csbj.2023.01.033
Daszkowska-Golec A, Mascher M, Zhang R:
Editorial: Applications of long-read sequencing in plant genomics and transcriptomics. Front. Plant Sci. 14 (2023) 1141429. https://dx.doi.org/10.3389/fpls.2023.1141429
El Hanafi S, Jiang Y, Kehel Z, Schulthess A W, Zhao Y, Mascher M, Haupt M, Himmelbach A, Stein N, Amri A, Reif J C:
Genomic predictions to leverage phenotypic data across genebanks. Front. Plant Sci. 14 (2023) 1227656. https://dx.doi.org/10.3389/fpls.2023.1227656
Feng J-W, Mascher M:
The story of wheat and its cousins. Nat. Plants 9 (2023) 377-378. https://dx.doi.org/10.1038/s41477-023-01365-5
Gemsa C:
Simulatios of the condensation and de-condensation of the barley chromatin during the cell cycle. (Bachelor Thesis) Halle/S., Martin-Luther-Universität Halle-Wittenberg, Naturwissenschaftliche Fakultät III Agrar- und Ernährungswissenschaften, Geowissenschaften und Informatik (2023)
Huang Y, Kamal R, Shanmugaraj N, Rutten T, Thirulogachandar V, Zhao S, Hoffie I, Hensel G, Rajaraman J, Moya Y A T, Hajirezaei M-R, Himmelbach A, Poursarebani N, Lundqvist U, Kumlehn J, Stein N, von Wirén N, Mascher M, Melzer M, Schnurbusch T:
A molecular framework for grain number determination in barley. Sci. Adv. 9 (2023) eadd0324. https://dx.doi.org/10.1126/sciadv.add0324
Jayakodi M, Golicz A A, Kreplak J, Fechete L I, Angra D, Bednář P, Bornhofen E, Zhang H, Boussageon R, Kaur S, Cheung K, Čížková J, Gundlach H, Hallab A, Imbert B, Keeble-Gagnère G, Koblížková A, Kobrlová L, Krejčí P, Mouritzen T W, Neumann P, Nadzieja M, Nielsen L K, Novák P, Orabi J, Padmarasu S, Robertson-Shersby-Harvie T, Robledillo L Á, Schiemann A, Tanskanen J, Törönen P, Warsame A O, Wittenberg A H J, Himmelbach A, Aubert G, Courty P-E, Doležel J, Holm L U, Janss L L, Khazaei H, Macas J, Mascher M, Smýkal P, Snowdon R J, Stein N, Stoddard F L, Stougaard J, Tayeh N, Torres A M, Usadel B, Schubert I, O'Sullivan D M, Schulman A H, Andersen S U:
The giant diploid faba genome unlocks variation in a global protein crop. Nature 615 (2023) 652-659. https://dx.doi.org/10.1038/s41586-023-05791-5
König P, Beier S, Mascher M, Stein N, Lange M, Scholz U:
DivBrowse—interactive visualization and exploratory data analysis of variant call matrices. GigaScience 12 (2023) giad025. https://dx.doi.org/10.1093/gigascience/giad025
Kubalová I, Câmara A S, Cápal P, Beseda T, Rouillard J-M, Krause Gina M, Holušová K, Toegelová H, Himmelbach A, Stein N, Houben A, Doležel J, Mascher M, Šimková H, Schubert V:
Helical coiling of metaphase chromatids. Nucleic Acids Res. 51 (2023) 2641-2654. https://dx.doi.org/10.1093/nar/gkad028
Kuo Y-T, Câmara A S, Schubert V, Neumann P, Macas J, Melzer M, Chen J, Fuchs J, Abel S, Klocke E, Huettel B, Himmelbach A, Demidov D, Dunemann F, Mascher M, Ishii T, Marques A, Houben A:
Holocentromeres can consist of merely a few megabase-sized satellite arrays. Nat. Commun. 14 (2023) 3502. https://dx.doi.org/10.1038/s41467-023-38922-7
Maniero R A, Koltun A, Vitti M, Factor B G, de Setta N, Câmara A S, Lima J E, Figueira A:
Identification and functional characterization of the sugarcane (Saccharum spp.) AMT2-type ammonium transporter ScAMT3;3 revealed a presumed role in shoot ammonium remobilization. Front. Plant Sci. 14 (2023) 1299025. https://dx.doi.org/10.3389/fpls.2023.1299025
Ost C, Cao H X, Nguyen T L, Himmelbach A, Mascher M, Stein N, Humbeck K:
Drought-stress-related reprogramming of gene expression in barley involves differential histone modifications at ABA-related genes. Int. J. Mol. Sci. 24 (2023) 12065. https://dx.doi.org/10.3390/ijms241512065
Shanmugaraj N, Rajaraman J, Kale S, Kamal R, Huang Y, Thirulogachandar V, Garibay-Hernandez A, Budhagatapalli N, Tandron Moya Y A, Hajirezaei M R, Rutten T, Hensel G, Melzer M, Kumlehn J, von Wirén N, Mock H-P, Schnurbusch T:
Multilayered regulation of developmentally programmed pre-anthesis tip degeneration of the barley inflorescence. Plant Cell 35 (2023) 3973-4001. https://dx.doi.org/10.1093/plcell/koad164
Acevedo-Garcia J, Walden K, Leissing F, Baumgarten K, Drwiega K, Kwaaitaal M, Reinstädler A, Freh M, Dong X, James G V, Baus L C, Mascher M, Stein N, Schneeberger K, Brocke-Ahmadinejad N, Kollmar M, Schulze-Lefert P, Panstruga R:
Barley Ror1 encodes a class XI myosin required for mlo-based broad-spectrum resistance to the fungal powdery mildew pathogen. Plant J. 112 (2022) 84-103. https://dx.doi.org/10.1111/tpj.15930
Avni R, Lux T, Minz-Dub A, Millet E, Sela H, Distelfeld A, Deek J, Yu G, Steuernagel B, Pozniak C, Ens J, Gundlach H, Mayer K F X, Himmelbach A, Stein N, Mascher M, Spannagl M, Wulff B B H, Sharon A:
Genome sequences of three Aegilops species of the section Sitopsis reveal phylogenetic relationships and provide resources for wheat improvement. Plant J. 110 (2022) 179-192. https://dx.doi.org/10.1111/tpj.15664
Chang C-W, Fridman E, Mascher M, Himmelbach A, Schmid K:
Physical geography, isolation by distance and environmental variables shape genomic variation of wild barley (Hordeum vulgare L. ssp. spontaneum) in the Southern Levant. Heredity 128 (2022) 107–119. https://dx.doi.org/10.1038/s41437-021-00494-x
Demidov D, Lermontova I, Moebes M, Kochevenko A, Fuchs J, Weiss O, Rutten T, Sorge E, Zuljan E, Giehl R F H, Mascher M, Somasundaram S, Conrad U, Houben A:
Haploid induction by nanobody targeted ubiquitin-proteasome-based degradation of EYFP-tagged CENH3 in Arabidopsis thaliana. J. Exp. Bot. 73 (2022) 7243–7254. https://dx.doi.org/10.1093/jxb/erac359
Dinh H X, Singh D, Gomez de la Cruz D, Hensel G, Kumlehn J, Mascher M, Stein N, Perovic D, Ayliffe M, Moscou M J, Park R F, Pourkheirandish M:
The barley leaf rust resistance gene Rph3 encodes a predicted membrane protein and is induced upon infection by avirulent pathotypes of Puccinia hordei. Nat. Commun. 13 (2022) 2386. https://dx.doi.org/10.1038/s41467-022-29840-1
Dreissig S, Mascher M:
Cherish your weeds. Mol. Plant 15 (2022) 396-397. https://dx.doi.org/10.1016/j.molp.2022.01.021
Fusi R, Rosignoli S, Lou H, Sangiorgi G, Bovina R, Pattem J K, Borkar A N, Lombardi M, Forestan C, Milner S G, Davis J L, Lale A, Kirschner G K, Swarup R, Tassinari A, Pandey B K, York L M, Atkinson B S, Sturrock C J, Mooney S J, Hochholdinger F, Tucker M R, Himmelbach A, Stein N, Mascher M, Nagel K A, de Gara L, Simmonds J, Uauy C, Tuberosa R, Lynch J P, Yakubov G E, Bennett M J, Bhosale R, Salvi S:
Root angle is controlled by EGT1 in cereal crops employing a novel anti-gravitropic mechanism. Proc. Natl. Acad. Sci. U.S.A. 119 (2022) e2201350119. https://dx.doi.org/10.1073/pnas.2201350119
Garg V, Dudchenko O, Wang J, Khan A W, Gupta S, Kaur P, Han K, Saxena R K, Kale S M, Pham M, Yu J, Chitikineni A, Zhang Z, Fan G, Lui C, Valluri V, Meng F, Bhandari A, Liu X, Yang T, Chen H, Valliyodan B, Roorkiwal M, Shi C, Yang H B, Durand N C, Pandey M K, Li G, Barmukh R, Wang X, Chen X, Lam H-M, Jiang H, Zong X, Liang X, Liu X, Liao B, Guo B, Jackson S, Nguyen H T, Zhuang W, Shubo W, Wang X, Aiden E L, Bennetzen J L, Varshney R K:
Chromosome-length genome assemblies of six legume species provide insights into genome organization, evolution, and agronomic traits for crop improvement. J. Adv. Res. 42 (2022) 315-329. https://dx.doi.org/10.1016/j.jare.2021.10.009
Gaurav K, Arora S, Silva P, Sanchez-Martin J, Horsnell R, Gao L, Brar G S, Widrig V, John Raupp W, Singh N, Wu S, Kale S M, Chinoy C, Nicholson P, Quiroz-Chavez J, Simmonds J, Hayta S, Smedley M A, Harwood W, Pearce S, Gilbert D, Kangara N, Gardener C, Forner-Martinez M, Liu J, Yu G, Boden S A, Pascucci A, Ghosh S, Hafeez A N, O'Hara T, Waites J, Cheema J, Steuernagel B, Patpour M, Justesen A F, Liu S, Rudd J C, Avni R, Sharon A, Steiner B, Kirana R P, Buerstmayr H, Mehrabi A A, Nasyrova F Y, Chayut N, Matny O, Steffenson B J, Sandhu N, Chhuneja P, Lagudah E, Elkot A F, Tyrrell S, Bian X, Davey R P, Simonsen M, Schauser L, Tiwari V K, Randy Kutcher H, Hucl P, Li A, Liu D-C, Mao L, Xu S, Brown-Guedira G, Faris J, Dvorak J, Luo M-C, Krasileva K, Lux T, Artmeier S, Mayer K F X, Uauy C, Mascher M, Bentley A R, Keller B, Poland J, Wulff B B H:
Population genomic analysis of Aegilops tauschii identifies targets for bread wheat improvement. Nat. Biotechnol. 40 (2022) 422–431. https://dx.doi.org/10.1038/s41587-021-01058-4
Guo Y, Himmelbach A, Weiss E, Stein N, Mascher M:
Six-rowed wild-growing barleys are hybrids of diverse origins. Plant J. 111 (2022) 849-858. https://dx.doi.org/10.1111/tpj.15861
Hinterberger V, Douchkov D, Lück S, Kale S, Mascher M, Stein N, Reif J C, Schulthess A W:
Mining for new sources of resistance to powdery mildew in genetic resources of winter wheat. Front. Plant Sci. 13 (2022) 836723. https://dx.doi.org/10.3389/fpls.2022.836723
Jiang C, Lei M, Guo Y, Gao G, Shi L, Jin Y, Cai Y, Himmelbach A, Zhou S, He Q, Yao X, Kan J, Haberer G, Duan F, Li L, Liu J, Zhang J, Spannagl M, Liu C, Stein N, Feng Z, Mascher M, Yang P:
A reference-guided TILLING by amplicon-seq platform supports forward and reverse genetics in barley. Plant Commun. 3 (2022) 100317. https://doi.org/10.1016/j.xplc.2022.100317
Kale S M, Schulthess A W, Padmarasu S, Boeven P H G, Schacht J, Himmelbach A, Steuernagel B, Wulff B B H, Reif J C, Stein N, Mascher M:
A catalogue of resistance gene homologs and a chromosome-scale reference sequence support resistance gene mapping in winter wheat. Plant Biotechnol. J. 20 (2022) 1730-1742. https://dx.doi.org/10.1111/pbi.13843
Kamal N, Lux T, Jayakodi M, Haberer G, Gundlach H, Mayer K F X, Mascher M, Spannagl M:
The barley and wheat pan-genomes. In: Edwards D (Ed.): Plant bioinformatics: methods and protocols. (Series: Methods in molecular biology, Vol. 2443) New York, NY: Humana (2022) ISBN 978-1-0716-2066-3, 147-159. https://dx.doi.org/10.1007/978-1-0716-2067-0_7
Kamal N, Tsardakas Renhuldt N, Bentzer J, Gundlach H, Haberer G, Juhász A, Lux T, Bose U, Tye-Din J A, Lang D, van Gessel N, Reski R, Fu Y-B, Spégel P, Ceplitis A, Himmelbach A, Waters A J, Bekele W A, Colgrave M L, Hansson M, Stein N, Mayer K F X, Jellen E N, Maughan P J, Tinker N A, Mascher M, Olsson O, Spannagl M, Sirijovski N:
The mosaic oat genome gives insights into a uniquely healthy cereal crop. Nature 606 (2022) 113-119. https://dx.doi.org/10.1038/s41586-022-04732-y
Koppolu R, Jiang G, Milner S G, Muqaddasi Q H, Rutten T, Himmelbach A, Guo Y, Stein N, Mascher M, Schnurbusch T:
The barley mutant multiflorus2.b reveals quantitative genetic variation for new spikelet architecture. Theor. Appl. Genet. 135 (2022) 571–590. https://dx.doi.org/10.1007/s00122-021-03986-w
Navrátilová P, Toegelová H, Tulpová Z, Kuo Y-T, Stein N, Doležel J, Houben A, Šimková H, Mascher M:
Prospects of telomere-to-telomere assembly in barley: analysis of sequence gaps in the MorexV3 reference genome. Plant Biotechnol. J. 20 (2022) 1373-1386. https://dx.doi.org/10.1111/pbi.13816
Püpke Marone M, Singh H C, Pozniak C J, Mascher M:
A technical guide to TRITEX, a computational pipeline for chromosome-scale sequence assembly of plant genomes. Plant Methods 18 (2022) 128. https://dx.doi.org/10.1186/s13007-022-00964-1
Rosignoli S, Cosenza F, Moscou M J, Civolani L, Musiani F, Forestan C, Milner S G, Savojardo C, Tuberosa R, Salvi S:
Cloning the barley nec3 disease lesion mimic mutant using complementation by sequencing. Plant Genome 15 (2022) e20187. https://dx.doi.org/10.1002/tpg2.20187
Sakkour A, Mascher M, Himmelbach A, Haberer G, Lux T, Spannagl M, Stein N, Kawamoto S, Sato K:
Chromosome-scale assembly of barley Cv. 'Haruna Nijo' as a resource for barley genetics. DNA Res. 29 (2022) dsac001. https://dx.doi.org/10.1093/dnares/dsac001
Schreiber M, Gao Y, Koch N, Fuchs J, Heckmann S, Himmelbach A, Börner A, Özkan H, Maurer A, Stein N, Mascher M, Dreissig S:
Recombination landscape divergence between populations is marked by larger low-recombining regions in domesticated rye. Mol. Biol. Evol. 39 (2022) msac131. https://dx.doi.org/10.1093/molbev/msac131
Schulthess A W, Kale S M, Liu F, Zhao Y, Philipp N, Rembe M, Jiang Y, Beukert U, Serfling A, Himmelbach A, Fuchs J, Oppermann M, Weise S, Boeven P H G, Schacht J, Longin C F H, Kollers S, Pfeiffer N, Korzun V, Lange M, Scholz U, Stein N, Mascher M, Reif J C:
Genomics-informed prebreeding unlocks the diversity in genebanks for wheat improvement. Nat. Genet. 54 (2022) 1544-1552. https://dx.doi.org/10.1038/s41588-022-01189-7
Schulthess A W, Kale S M, Zhao Y, Gogna A, Rembe M, Philipp N, Liu F, Beukert U, Serfling A, Himmelbach A, Oppermann M, Weise S, Boeven P H G, Schacht J, Longin C F H, Kollers S, Pfeiffer N, Korzun V, Fiebig A, Schüler D, Lange M, Scholz U, Stein N, Mascher M, Reif J C:
Large-scale genotyping and phenotyping of a worldwide winter wheat genebank for its use in pre-breeding. Sci. Data 9 (2022) 784. https://dx.doi.org/10.1038/s41597-022-01891-5
Tinker N A, Wight C P, Bekele W A, Yan W, Jellen E N, Renhuldt N T, Sirijovski N, Lux T, Spannagl M, Mascher M:
Genome analysis in Avena sativa reveals hidden breeding barriers and opportunities for oat improvement. Commun. Biol. 5 (2022) 474. https://dx.doi.org/10.1038/s42003-022-03256-5
Wang Y, Habekuss A, Jayakodi M, Mascher M, Snowdon R J, Stahl A, Fuss J, Ordon F, Perovic D:
High-resolution mapping of Barley mild mosaic virus resistance gene rym15. Front. Plant Sci. 13 (2022) 908170. https://dx.doi.org/10.3389/fpls.2022.908170
Yu G, Matny O, Champouret N, Steuernagel B, Moscou M J, Hernández-Pinzón I, Green P, Hayta S, Smedley M, Harwood W, Kangara N, Yue Y, Gardener C, Banfield M J, Olivera P D, Welchin C, Simmons J, Millet E, Minz-Dub A, Ronen M, Avni R, Sharon A, Patpour M, Justesen A F, Jayakodi M, Himmelbach A, Stein N, Wu S, Poland J, Ens J, Pozniak C, Karafiátová M, Molnár I, Doležel J, Ward E R, Reuber T L, Jones J D G, Mascher M, Steffenson B J, Wulff B B H:
Aegilops sharonensis genome-assisted identification of stem rust resistance gene Sr62. Nat. Commun. 13 (2022) 1607. https://dx.doi.org/10.1038/s41467-022-29132-8
Zhang H, Mascher M, Abbo S, Jayakodi M:
Advancing grain legumes domestication and evolution studies with genomics. Plant Cell Physiol. 63 (2022) 1540–1553. https://dx.doi.org/10.1093/pcp/pcac062
Câmara A S, Schubert V, Mascher M, Houben A:
A simple model explains the cell cycle-dependent assembly of centromeric nucleosomes in holocentric species. Nucleic Acids Res. 49 (2021) 9053–9065. https://dx.doi.org/10.1093/nar/gkab648
Chen C, Jost M, Clark B, Martin M, Matny O, Steffenson B J, Franckowiak J D, Mascher M, Singh D, Perovic D, Richardson T, Periyannan S, Lagudah E S, Park R F, Dracatos P M:
BED-domain-containing NLR from wild barley confers resistance to leaf rust. Plant Biotechnol. J. 19 (2021) 1206-1215. https://dx.doi.org/10.1111/pbi.13542
Gao L, Koo D-H, Juliana P, Rife T, Singh D, Lemes da Silva C, Lux T, Dorn K M, Clinesmith M, Silva P, Wang X, Spannagl M, Monat C, Friebe B, Steuernagel B, Muehlbauer G J, Walkowiak S, Pozniak C, Singh R, Stein N, Mascher M, Fritz A, Poland J:
The Aegilops ventricosa 2NvS segment in bread wheat: cytology, genomics and breeding. Theor. Appl. Genet. 134 (2021) 529–542. https://dx.doi.org/10.1007/s00122-020-03712-y
Halstead-Nussloch G, Tanaka T, Copetti D, Paape T, Kobayashi F, Hatakeyama M, Kanamori H, Wu J, Mascher M, Kawaura K, Shimizu K K, Handa H:
Multiple wheat genomes reveal novel Gli-2 sublocus location and variation of celiac disease epitopes in duplicated alpha-gliadin genes. Front. Plant Sci. 12 (2021) 715985. https://dx.doi.org/10.3389/fpls.2021.715985
Hewitt T, Müller M C, Molnár I, Mascher M, Holušová K, Šimková H, Kunz L, Zhang J, Li J, Bhatt D, Sharma R, Schudel S, Yu G, Steuernagel B, Periyannan S, Wulff B, Ayliffe M, McIntosh R, Keller B, Lagudah E, Zhang P:
A highly differentiated region of wheat chromosome 7AL encodes a Pm1a immune receptor that recognises its corresponding AvrPm1a effector from Blumeria graminis. New Phytol. 229 (2021) 2812-2826. https://dx.doi.org/10.1111/nph.17075
Jayakodi M, Schreiber M, Stein N, Mascher M:
Building pan-genome infrastructures for crop plants and their use in association genetics. DNA Res. 28 (2021) dsaa030. https://dx.doi.org/10.1093/dnares/dsaa030
Kirschner G K, Rosignoli S, Guo L, Vardanega I, Imani J, Altmüller J, Milner S G, Balzano R, Nagel K A, Pflugfelder D, Forestan C, Bovina R, Koller R, Stöcker T G, Mascher M, Simmonds J, Uauy C, Schoof H, Tuberosa R, Salvi S, Hochholdinger F:
ENHANCED GRAVITROPISM 2 encodes a STERILE ALPHA MOTIF-containing protein that controls root growth angle in barley and wheat. Proc. Natl. Acad. Sci. U.S.A. 118 (2021) e2101526118. https://dx.doi.org/10.1073/pnas.2101526118
Lange M, Alako B T F, Cochrane G, Ghaffar M, Mascher M, Habekost P-K, Hillebrand U, Scholz U, Schorch F, Freitag J, Scholz A H:
Quantitative monitoring of nucleotide sequence data from genetic resources in context of their citation in the scientific literature. GigaScience 10 (2021) giab084. https://dx.doi.org/10.1093/gigascience/giab084
Mascher M, Jayakodi M, Stein N:
The reinvention of potato. Cell Res. 31 (2021) 1144–1145. https://dx.doi.org/10.1038/s41422-021-00542-5
Mascher M, Wicker T, Jenkins J, Plott C, Lux T, Koh C S, Ens J, Gundlach H, Boston L B, Tulpová Z, Holden S, Hernández-Pinzón I, Scholz U, Mayer K F X, Spannagl M, Pozniak C J, Sharpe A G, Šimková H, Moscou M J, Grimwood J, Schmutz J, Stein N:
Long-read sequence assembly: a technical evaluation in barley. Plant Cell 33 (2021) 1888-1906. https://dx.doi.org/10.1093/plcell/koab077
Petroll R, Schreiber M, Finke H, Cock J M, Gould S B, Rensing S A:
Signatures of transcription factor evolution and the secondary gain of red algae complexity. Genes 12 (2021) 1055. https://dx.doi.org/10.3390/genes12071055
Rabanus-Wallace M T, Hackauf B, Mascher M, Lux T, Wicker T, Gundlach H, Baez M, Houben A, Mayer K F X, Guo L, Poland J, Pozniak C J, Walkowiak S, Melonek J, Praz C R, Schreiber M, Budak H, Heuberger M, Steuernagel B, Wulff B, Börner A, Byrns B, Čížková J, Fowler D B, Fritz A, Himmelbach A, Kaithakottil G, Keilwagen J, Keller B, Konkin D, Larsen J, Li Q, Myśków B, Padmarasu S, Rawat N, Sesiz U, Biyiklioglu-Kaya S, Sharpe A, Šimková H, Small I, Swarbreck D, Toegelová H, Tsvetkova N, Voylokov A V, Vrána J, Bauer E, Bolibok-Bragoszewska H, Doležel J, Hall A, Jia J, Korzun V, Laroche A, Ma X-F, Ordon F, Özkan H, Rakoczy-Trojanowska M, Scholz U, Schulman A H, Siekmann D, Stojalowski S, Tiwari V K, Spannagl M, Stein N:
Chromosome-scale genome assembly provides insights into rye biology, evolution and agronomic potential. Nat. Genet. 53 (2021) 564-573. https://dx.doi.org/10.1038/s41588-021-00807-0
Sato K, Abe F, Mascher M, Haberer G, Gundlach H, Spannagl M, Shirasawa K, Isobe S:
Chromosome-scale genome assembly of the transformation-amenable common wheat cultivar 'Fielder'. DNA Res. 28 (2021) dsab008. https://dx.doi.org/10.1093/dnares/dsab008
Sato K, Mascher M, Himmelbach A, Haberer G, Spannagl M, Stein N:
Chromosome-scale assembly of wild barley accession ‘OUH602’. G3 Genes Genom. Genet. 11 (2021) jkab244. https://doi.org/10.1093/g3journal/jkab244
Schreiber M, Özkan H, Komatsuda T, Mascher M:
Evolution and domestication of rye. In: Rabanus-Wallace M T, Stein N (Eds.): The rye genome. (Series: Compendium of plant genomes) Cham: Springer (2021) ISBN 978-3-030-83383-1, 85-100. https://dx.doi.org/10.1007/978-3-030-83383-1_6
Shimizu K K, Copetti D, Okada M, Wicker T, Tameshige T, Hatakeyama M, Shimizu-Inatsugi R, Aquino C, Nishimura K, Kobayashi F, Murata K, Kuo T, Delorean E, Poland J, Haberer G, Spannagl M, Mayer K F X, Gutierrez-Gonzalez J, Muehlbauer G J, Monat C, Himmelbach A, Padmarasu S, Mascher M, Walkowiak S, Nakazaki T, Ban T, Kawaura K, Tsuji H, Pozniak C, Stein N, Sese J, Nasuda S, Handa H:
De novo genome assembly of the Japanese wheat cultivar Norin 61 highlights functional variation in flowering time and Fusarium resistance genes in East Asian genotypes. Plant Cell Physiol. 62 (2021) 8-27. https://dx.doi.org/10.1093/pcp/pcaa152
Thiel J, Koppolu R, Trautewig C, Hertig C, Kale S M, Erbe S, Mascher M, Himmelbach A, Rutten T, Esteban E, Pasha A, Kumlehn J, Provart N J, Vanderauwera S, Frohberg C, Schnurbusch T:
Transcriptional landscapes of floral meristems in barley. Sci. Adv. 7 (2021) eabf0832. https://dx.doi.org/10.1126/sciadv.abf0832
Tripodi P, Rabanus-Wallace M T, Barchi L, Kale S, Esposito S, Acquadro A, Schafleitner R, van Zonneveld M, Prohens J, Diez M J, Börner A, Salinier J, Caromel B, Bovy A, Boyaci F, Pasev G, Brandt R, Himmelbach A, Portis E, Finkers R, Lanteri S, Paran I, Lefebvre V, Giuliano G, Stein N:
Global range expansion history of pepper (Capsicum spp.) revealed by over 10,000 genebank accessions. Proc. Natl. Acad. Sci. U.S.A. 118 (2021) e2104315118. https://dx.doi.org/10.1073/pnas.2104315118
Wolde G M, Schreiber M, Trautewig C, Himmelbach A, Sakuma S, Mascher M, Schnurbusch T:
Genome-wide identification of loci modifying spike-branching in tetraploid wheat. Theor. Appl. Genet. 134 (2021) 1925–1943. https://dx.doi.org/10.1007/s00122-020-03743-5
Xu W, Tucker J R, Bekele W A, You F M, Fu Y-B, Khanal R, Yao Z, Singh J, Boyle B, Beattie A D, Belzile F, Mascher M, Tinker N A, Badea A:
Genome assembly of the Canadian two-row malting barley cultivar AAC synergy. G3 Genes Genom. Genet. 11 (2021) jkab031. https://dx.doi.org/10.1093/g3journal/jkab031
Žegarac A, Winkelbach L, Blöcher J, Diekmann Y, Krečković Gavrilović M, Porčić M, Stojković B, Milašinović L, Schreiber M, Wegmann D, Veeramah K R, Stefanović S, Burger J:
Ancient genomes provide insights into family structure and the heredity of social status in the early Bronze Age of southeastern Europe. Sci. Rep. 11 (2021) 10072. https://dx.doi.org/10.1038/s41598-021-89090-x
Zhu T, Wang L, Rimbert H, Rodriguez J C, Deal K R, De Oliveira R, Choulet F, Keeble-Gagnère G, Tibbits J, Rogers J, Eversole K, Appels R, Gu Y Q, Mascher M, Dvorak J, Luo M-C:
Optical maps refine the bread wheat Triticum aestivum cv Chinese Spring genome assembly. Plant J. 107 (2021) 303-314. https://doi.org/10.1111/tpj.15289
Dreissig S, Fuchs J, Himmelbach A, Mascher M, Houben A:
Quantification of recombination rate and segregation distortion by genotyping and sequencing of single pollen nuclei. In: Pradillo M, Heckmann S (Eds.): Plant Meiosis: methods and protocols. (Series: Methods in molecular biology, Vol. 2061) New York, NY: Humana Press (2020) 978-1-4939-9817-3, 281-300. https://dx.doi.org/10.1007/978-1-4939-9818-0_20
Haas M, Himmelbach A, Mascher M:
The contribution of cis- and trans-acting variants to gene regulation in wild and domesticated barley under cold stress and control conditions. J. Exp. Bot. 71 (2020) 2573-2584. https://dx.doi.org/10.1093/jxb/eraa036
Jayakodi M, Padmarasu S, Haberer G, Bonthala V S, Gundlach H, Monat C, Lux T, Kamal N, Lang D, Himmelbach A, Ens J, Zhang X Q, Angessa T T, Zhou G, Tan C, Hill C, Wang P, Schreiber M, Boston L B, Plott C, Jenkins J, Guo Y, Fiebig A, Budak H, Xu D, Zhang J, Wang C, Grimwood J, Schmutz J, Guo G, Zhang G, Mochida K, Hirayama T, Sato K, Chalmers K J, Langridge P, Waugh R, Pozniak C J, Scholz U, Mayer K F X, Spannagl M, Li C, Mascher M, Stein N:
The barley pan-genome reveals the hidden legacy of mutation breeding. Nature 588 (2020) 284-289. https://dx.doi.org/10.1038/s41586-020-2947-8
König P, Beier S, Basterrechea M, Schüler D, Arend D, Mascher M, Stein N, Scholz U, Lange M:
BRIDGE – a visual analytics web tool for barley genebank genomics. Front. Plant Sci. 11 (2020) 701. https://dx.doi.org/10.3389/fpls.2020.00701
Saxena R K, Kale S, Mir R R, Mallikarjuna N, Yadav P, Das R R, Molla J, Sonnappa M, Ghanta A, Narasimhan Y, Rathore A, Kumar C V S, Varshney R K:
Genotyping-by-sequencing and multilocation evaluation of two interspecific backcross populations identify QTLs for yield-related traits in pigeonpea. Theor. Appl. Genet. 133 (2020) 737–749. https://dx.doi.org/10.1007/s00122-019-03504-z
Schreiber M, Mascher M, Wright J, Padmarasu S, Himmelbach A, Heavens D, Milne L, Clavijo B J, Stein N, Waugh R:
A genome assembly of the barley 'transformation reference' cultivar Golden Promise. G3-Genes Genom. Genet. 10 (2020) 1823-1827. https://dx.doi.org/10.1534/g3.119.401010
Sinha P, Singh V K, Saxena R K, Kale S M, Li Y, Garg V, Tang M, Khan A W, Kim K D, Chitikineni A, Saxena K B, Sameer Kumar C V, Liu X, Xu X, Jackson S, Powell W, Nevo E, Searle I R, Lodha M, Varshney R K:
Genome-wide analysis of epigenetic and transcriptional changes associated with heterosis in pigeonpea. Plant Biotechnol. J. 18 (2020) 1697-1710. https://dx.doi.org/10.1111/pbi.13333
Tikhenko N, Alqudah A M, Borisjuk L, Ortleb S, Rutten T, Wu D D, Nagel M, Himmelbach A, Mascher M, Röder M, Ganal M, Sehmisch S, Houben A, Börner A:
DEFECTIVE ENDOSPERM-D1 (Dee-D1) is crucial for endosperm development in hexaploid wheat. Commun. Biol. 3 (2020) 791. https://doi.org/10.1038/s42003-020-01509-9
Walkowiak S, Gao L, Monat C, Haberer G, Kassa M T, Brinton J, Ramirez-Gonzalez R H, Kolodziej M C, Delorean E, Thambugala D, Klymiuk V, Byrns B, Gundlach H, Bandi V, Siri J N, Nilsen K, Aquino C, Himmelbach A, Copetti D, Ban T, Venturini L, Bevan M, Clavijo B, Koo D H, Ens J, Wiebe K, N'Diaye A, Fritz A K, Gutwin C, Fiebig A, Fosker C, Fu B X, Accinelli G G, Gardner K A, Fradgley N, Gutierrez-Gonzalez J, Halstead-Nussloch G, Hatakeyama M, Koh C S, Deek J, Costamagna A C, Fobert P, Heavens D, Kanamori H, Kawaura K, Kobayashi F, Krasileva K, Kuo T, McKenzie N, Murata K, Nabeka Y, Paape T, Padmarasu S, Percival-Alwyn L, Kagale S, Scholz U, Sese J, Juliana P, Singh R, Shimizu-Inatsugi R, Swarbreck D, Cockram J, Budak H, Tameshige T, Tanaka T, Tsuji H, Wright J, Wu J, Steuernagel B, Small I, Cloutier S, Keeble-Gagnere G, Muehlbauer G, Tibbets J, Nasuda S, Melonek J, Hucl P J, Sharpe A G, Clark M, Legg E, Bharti A, Langridge P, Hall A, Uauy C, Mascher M, Krattinger S G, Handa H, Shimizu K K, Distelfeld A, Chalmers K, Keller B, Mayer K F X, Poland J, Stein N, McCartney C A, Spannagl M, Wicker T, Pozniak C J:
Multiple wheat genomes reveal global variation in modern breeding. Nature 588 (2020) 277-283. https://dx.doi.org/10.1038/s41586-020-2961-x
Youssef H M, Allam M, Boussora F, Himmelbach A, Milner S G, Mascher M, Schnurbusch T:
Dissecting the genetic basis of lateral and central spikelet development and grain traits in intermedium-spike barley (Hordeum vulgare convar. intermedium). Plants 9 (2020) 1655. https://dx.doi.org/10.3390/plants9121655
Börner A, Alqudah A, Alomari D, Berrueta W, Cardelli M, Castro A, Castro A, Chesnokov Y, del Río J, Eggert K, Giménez D, Jayakodi M, Kartseva T, Lohwasser U, Lori G, Malbrán I, Misheva S, Muqaddasi Q, Nagel M, Röder M, Saldúa L, Schierenbeck M, Shamanin V, Simón M, Tarawneh R, Uranga J, von Wirén N, Yanniccari M, Zaynali Nezhad K:
Items from Germany. Ann. Wheat Newsl. 65 (2019) 12-16.
Bustos-Korts D, Dawson I K, Russell J, Tondelli A, Guerra D, Ferrandi C, Strozzi F, Nicolazzi E L, Molnar-Lang M, Ozkan H, Megyeri M, Miko P, Cakir E, Yakisir E, Trabanco N, Delbono S, Kyriakidis S, Booth A, Cammarano D, Mascher M, Werner P, Cattivelli L, Rossini L, Stein N, Kilian B, Waugh R, van Eeuwijk F A:
Exome sequences and multi-environment field trials elucidate the genetic basis of adaptation in barley. Plant J. 99 (2019) 1172-1191. https://dx.doi.org/10.1111/tpj.14414
Darrier B, Russell J, Milner S G, Hedley P E, Shaw P D, Macaulay M, Ramsay L D, Halpin C, Mascher M, Fleury D L, Langridge P, Stein N, Waugh R:
A comparison of mainstream genotyping platforms for the evaluation and use of barley genetic resources. Front. Plant Sci. 10 (2019) 544. https://dx.doi.org/10.3389/fpls.2019.00544
Dreissig S, Mascher M, Heckmann S:
Variation in recombination rate is shaped by domestication and environmental conditions in barley. Mol. Biol. Evol. 36 (2019) 2029–2039. https://dx.doi.org/10.1093/molbev/msz141
Gutierrez-Gonzalez J J, Mascher M, Poland J, Muehlbauer G J:
Dense genotyping-by-sequencing linkage maps of two Synthetic W7984xOpata reference populations provide insights into wheat structural diversity. Sci. Rep. 9 (2019) 1793. https://dx.doi.org/10.1038/s41598-018-38111-3
Haas M, Mascher M:
Use of the secondary gene pool of barley in breeding improved varieties. In: Ordon F, Friedt W (Eds.): Advances in breeding techniques for cereal crops. (Series: Burleigh dodds series in agricultural science, Vol. 60) Cambridge, UK: Burleigh Dodds Science Pub LTD (2019) ISBN 978-1-78676-244-3, https://dx.doi.org/10.19103/AS.2019.0051.02
Haas M, Schreiber M, Mascher M:
Domestication and crop evolution of wheat and barley: Genes, genomics, and future directions. J. Integr. Plant Biol. 61 (2019) 204-225. https://dx.doi.org/10.1111/jipb.12737
Heuermann M C, Rosso M G, Mascher M, Brandt R, Tschiersch H, Altschmied L, Altmann T:
Combining next-generation sequencing and progeny testing for rapid identification of induced recessive and dominant mutations in maize M2 individuals. Plant J. 100 (2019) 851-862. https://dx.doi.org/10.1111/tpj.14431
Hoseinzadeh H, Zhou R, Mascher M, Himmelbach A, Niks R E, Schweizer P, Stein N:
High resolution genetic and physical mapping of a major powdery mildew resistance locus in barley. Front. Plant Sci. 10 (2019) 146. https://dx.doi.org/10.3389/fpls.2019.00146
Jayakodi M, Schreiber M, Mascher M:
Sweet genes in melon and watermelon. Nat. Genet. 51 (2019) 1572-1573. https://dx.doi.org/10.1038/s41588-019-0529-1
Li M, Hensel G, Mascher M, Melzer M, Budhagatapalli N, Rutten T, Himmelbach A, Beier S, Korzun V, Kumlehn J, Boerner T, Stein N:
Leaf variegation and impaired chloroplast development caused by a truncated CCT domain gene in albostrians barley. Plant Cell 31 (2019) 1430-1445. https://dx.doi.org/10.1105/tpc.19.00132
Maccaferri M, Harris N S, Twardziok S O, Pasam R K, Gundlach H, Spannagl M, Ormanbekova D, Lux T, Prade V M, Milner S G, Himmelbach A, Mascher M, Bagnaresi P, Faccioli P, Cozzi P, Lauria M, Lazzari B, Stella A, Manconi A, Gnocchi M, Moscatelli M, Avni R, Deek J, Biyiklioglu S, Frascaroli E, Corneti S, Salvi S, Sonnante G, Desiderio F, Mare C, Crosatti C, Mica E, Ozkan H, Kilian B, De Vita P, Marone D, Joukhadar R, Mazzucotelli E, Nigro D, Gadaleta A, Chao S, Faris J D, Melo A T O, Pumphrey M, Pecchioni N, Milanesi L, Wiebe K, Ens J, MacLachlan R P, Clarke J M, Sharpe A G, Koh C S, Liang K Y H, Taylor G J, Knox R, Budak H, Mastrangelo A M, Xu S S, Stein N, Hale I, Distelfeld A, Hayden M J, Tuberosa R, Walkowiak S, Mayer K F X, Ceriotti A, Pozniak C J, Cattivelli L:
Durum wheat genome highlights past domestication signatures and future improvement targets. Nat. Genet. 51 (2019) 885-895. https://dx.doi.org/10.1038/s41588-019-0381-3
Mascher M, Schreiber M, Scholz U, Graner A, Reif J C, Stein N:
Genebank genomics bridges the gap between the conservation of crop diversity and plant breeding. Nat. Genet. 51 (2019) 1076-1081. https://dx.doi.org/10.1038/s41588-019-0443-6
Milne R J, Dibley K E, Schnippenkoetter W H, Mascher M, Lui A C, Wang L, Lo C, Ashton A R, Ryan P R, Lagudah E:
The wheat Lr67 gene of the Sugar Transport Protein 13 family confers multipathogen resistance in barley. Plant Physiol. 179 (2019) 1285-1297. https://dx.doi.org/10.1104/pp.18.00945
Milner S G, Jost M, Taketa S, Mazón E R, Himmelbach A, Oppermann M, Weise S, Knüpffer H, Basterrechea M, König P, Schüler D, Sharma R, Pasam R K, Rutten T, Guo G, Xu D, Zhang J, Herren G, Müller T, Krattinger S G, Keller B, Jiang Y, González M Y, Zhao Y, Habekuß A, Färber S, Ordon F, Lange M, Börner A, Graner A, Reif J C, Scholz U, Mascher M, Stein N:
Genebank genomics highlights the diversity of a global barley collection. Nat. Genet. 51 (2019) 319-326. https://doi.org/10.1038/s41588-018-0266-x
Monat C, Padmarasu S, Lux T, Wicker T, Gundlach H, Himmelbach A, Ens J, Li C, Muehlbauer G J, Schulman A H, Waugh R, Braumann I, Pozniak C, Scholz U, Mayer K F X, Spannagl M, Stein N, Mascher M:
TRITEX: chromosome-scale sequence assembly of Triticeae genomes with open-source tools. Genome Biol. 20 (2019) 284. https://dx.doi.org/10.1186/s13059-019-1899-5
Monat C, Schreiber M, Stein N, Mascher M:
Prospects of pan-genomics in barley. Theor. Appl. Genet. 132 (2019) 785–796. https://dx.doi.org/10.1007/s00122-018-3234-z
Muqaddasi Q H, Jayakodi M, Börner A, Röder M S:
Identification of consistent QTL with large effect on anther extrusion in doubled haploid populations developed from spring wheat accessions in German Federal ex situ Genebank. Theor. Appl. Genet. 132 (2019) 3035–3045. https://dx.doi.org/10.1007/s00122-019-03404-2
Padmarasu S, Himmelbach A, Mascher M, Stein N:
In situ Hi-C for plants: an improved method to detect long-range chromatin interactions. In: Chekanova J, Wang H-L (Eds.): Plant long non-coding RNAs: methods and protocols. (Series: Methods in molecular biology, Vol. 1933) New York, NY: Humana Press (2019) ISBN 978-1-4939-9044-3, 441-472. https://doi.org/10.1007/978-1-4939-9045-0_28
Radchuk V, Sharma R, Potokina E, Radchuk R, Weier D, Munz E, Schreiber M, Mascher M, Stein N, Wicker T, Kilian B, Borisjuk L:
The highly divergent Jekyll genes, required for sexual reproduction, are lineage specific for the related grass tribes Triticeae and Bromeae. Plant J. 98 (2019) 961-974. https://dx.doi.org/10.1111/tpj.14363
Sakuma S, Golan G, Guo Z, Ogawa T, Tagiri A, Sugimoto K, Bernhardt N, Brassac J, Mascher M, Hensel G, Ohnishi S, Jinno H, Yamashita Y, Ayalon I, Peleg Z, Schnurbusch T, Komatsuda T:
Unleashing floret fertility in wheat through the mutation of a homeobox gene. Proc. Natl. Acad. Sci. U.S.A. 116 (2019) 5182-5187. https://dx.doi.org/10.1073/pnas.1815465116
Schreiber M:
Ein Blick in die Kulturgeschichte von Roggen, Gerste & Weizen. (PhD Thesis) Mainz, Johannes Gutenberg-Universität Mainz, Fachbereich Anthropologie (2019) 399 pp.
Schreiber M, Himmelbach A, Börner A, Mascher M:
Genetic diversity and relationship of domesticated rye and its wild relatives as revealed through genotyping-by-sequencing. Evol. Appl. 12 (2019) 66-77. https://dx.doi.org/10.1111/eva.12624
Wolde G M, Mascher M, Schnurbusch T:
Genetic modification of spikelet arrangement in wheat increases grain number without significantly affecting grain weight. Mol. Genet. Genomics 294 (2019) 457–468. https://dx.doi.org/10.1007/s00438-018-1523-5
Wolde G M, Trautewig C, Mascher M, Schnurbusch T:
Genetic insights into morphometric inflorescence traits of wheat. Theor. Appl. Genet. 132 (2019) 1661-1676. https://dx.doi.org/10.1007/s00122-019-03305-4
Brandt R, Mascher M, Thiel J:
Laser-capture microdissection-based RNA-seq of barley grain tissues. In: Murray G I (Ed.): Laser Capture Microdissection: Methods and Protocols. (Series: Methods in molecular biology, Vol. 1723) New York, NY: Humana Press (2018) ISBN 978-1-4939-7557-0, 397-409. https://doi.org/10.1007/978-1-4939-7558-7_23
Himmelbach A, Ruban A, Walde I, Šimková H, Doležel J, Hastie A, Stein N, Mascher M:
Discovery of multi-megabase polymorphic inversions by chromosome conformation capture sequencing in large-genome plant species. Plant J. 96 (2018) 1309-1316. https://dx.doi.org/10.1111/tpj.14109
Himmelbach A, Walde I, Mascher M, Stein N:
Tethered chromosome conformation capture sequencing in Triticeae: a valuable tool for genome assembly. Bio-protocol 8 (2018) e2955. https://dx.doi.org/10.21769/BioProtoc.2955
Mascher M, Sato K, Steffenson B:
Genomics approaches to mining barley germplasm collections. In: Stein N, Muehlbauer G J (Eds.): The Barley Genome, 1st ed. (Series: Kole, C (Ed.): Compendium of Plant Genomes) Cham: Springer (2018) ISBN 978-3-319-92528-8, 155-169. https://dx.doi.org/10.1007/978-3-319-92528-8_11
Muñoz-Amatriaín M, Mascher M:
Sequence diversity and structural variation. In: Stein N, Muehlbauer G J (Eds.): The Barley Genome, 1st ed. (Series: Kole, C (Ed.): Compendium of Plant Genomes) Cham: Springer (2018) ISBN 978-3-319-92528-8, 109-122. https://dx.doi.org/10.1007/978-3-319-92528-8_8
Rajaraman J, Douchkov D, Lück S, Hensel G, Nowara D, Pogoda M, Rutten T, Meitzel T, Brassac J, Höfle C, Hückelhoven R, Klinkenberg J, Trujillo M, Bauer E, Schmutzer T, Himmelbach A, Mascher M, Lazzari B, Stein N, Kumlehn J, Schweizer P:
Evolutionarily conserved partial gene duplication in the Triticeae tribe of grasses confers pathogen resistance. Genome Biol. 19 (2018) 116. https://dx.doi.org/10.1186/s13059-018-1472-7
Romero C C T, Vermeulen J P, Vels A, Himmelbach A, Mascher M, Niks R E:
Mapping resistance to powdery mildew in barley reveals a large-effect nonhost resistance QTL. Theor. Appl. Genet. 131 (2018) 1031–1045. https://dx.doi.org/10.1007/s00122-018-3055-0
Schreiber M, Stein N, Mascher M:
Genomic approaches for studying crop evolution. Genome Biol. 19 (2018) 140. https://dx.doi.org/10.1186/s13059-018-1528-8
Stein N, Mascher M:
Barley genome sequencing amd assembly - a first version reference sequence. In: Stein N, Muehlbauer G J (Eds.): The Barley Genome, 1st ed. (Series: Kole, C (Ed.): Compendium of Plant Genomes) Cham: Springer (2018) ISBN 978-3-319-92528-8, 57-71. https://dx.doi.org/10.1007/978-3-319-92528-8_5
The International Wheat Genome Sequencing Consortium (IWGSC; IPK authors: Mascher M, Zhou, R., Himmelbach, A. & co-corresponding author: Stein, N.):
Shifting the limits in wheat research and breeding using a fully annotated reference genome. Science 361 (2018) eaar7191. https://dx.doi.org/10.1126/science.aar7191
Zeng X, Guo Y, Xu Q, Mascher M, Guo G, Li S, Mao L, Liu Q, Xia Z, Zhou J, Yuan H, Tai S, Wang Y, Wei Z, Song L, Zha S, Li S, Tang Y, Bai L, Zhuang Z, He W, Zhao S, Fang X, Gao Q, Yin Y, Wang J, Yang H, Zhang J, Henry R J, Stein N, Tashi N:
Origin and evolution of qingke barley in Tibet. Nat. Commun. 9 (2018) 5433. https://dx.doi.org/10.1038/s41467-018-07920-5
Avni R, Nave M, Barad O, Baruch K, Twardziok S O, Gundlach H, Hale I, Mascher M, Spannagl M, Wiebe K, Jordan K W, Golan G, Deek J, Ben-Zvi B, Ben-Zvi G, Himmelbach A, MacLachlan R P, Sharpe A G, Fritz A, Ben-David R, Budak H, Fahima T, Korol A, Faris J D, Hernandez A, Mikel M A, Levy A A, Steffenson B, Maccaferri M, Tuberosa R, Cattivelli L, Faccioli P, Ceriotti A, Kashkush K, Pourkheirandish M, Komatsuda T, Eilam T, Sela H, Sharon A, Ohad N, Chamovitz D A, Mayer K F X, Stein N, Ronen G, Peleg Z, Pozniak C J, Akhunov E D, Distelfeld A:
Wild emmer genome architecture and diversity elucidate wheat evolution and domestication. Science 357 (2017) 93-97. https://dx.doi.org/10.1126/science.aan0032
Bauer E, Schmutzer T, Barilar I, Mascher M, Gundlach H, Martis M M, Twardziok S O, Hackauf B, Gordillo A, Wilde P, Schmidt M, Korzun V, Mayer K F X, Schmid K, Schön C-C, Scholz U:
Towards a whole-genome sequence for rye (Secale cereale L.). Plant J. 89 (2017) 853-869. https://dx.doi.org/10.1111/tpj.13436
Beier S, Himmelbach A, Colmsee C, Zhang X-Q, Barrero R A, Zhang Q, Li L, Bayer M, Bolser D, Taudien S, Groth M, Felder M, Hastie A, Šimková H, Staňková H, Vrána J, Chan S, Muñoz-Amatriaín M, Ounit R, Wanamaker S, Schmutzer T, Aliyeva-Schnorr L, Grasso S, Tanskanen J, Sampath D, Heavens D, Cao S, Chapman B, Dai F, Han Y, Li H, Li X, Lin C, McCooke J K, Tan C, Wang S, Yin S, Zhou G, Poland J A, Bellgard M I, Houben A, Doležel J, Ayling S, Lonardi S, Langridge P, Muehlbauer G J, Kersey P, Clark M D, Caccamo M, Schulman A H, Platzer M, Close T J, Hansson M, Zhang G, Braumann I, Li C, Waugh R, Scholz U, Stein N, Mascher M:
Construction of a map-based reference genome sequence for barley, Hordeum vulgare L. Sci. Data 4 (2017) 170044. https://dx.doi.org/10.1038/sdata.2017.44
Beier S, Thiel T, Münch T, Scholz U, Mascher M:
MISA-web: a web server for microsatellite prediction. Bioinformatics 33 (2017) 2583–2585. https://dx.doi.org/10.1093/bioinformatics/btx198
Braatz J, Harloff H-J, Mascher M, Stein N, Himmelbach A, Jung C:
CRISPR-Cas9 targeted mutagenesis leads to simultaneous modification of different homoeologous gene copies in polyploid oilseed rape (Brassica napus). Plant Physiol. 174 (2017) 935-942. https://dx.doi.org/10.1104/pp.17.00426
Dreissig S, Fuchs J, Himmelbach A, Mascher M, Houben A:
Sequencing of single pollen nuclei reveals meiotic recombination events at megabase resolution and circumvents segregation distortion caused by postmeiotic processes. Front. Plant Sci. 8 (2017) 1620. https://dx.doi.org/10.3389/fpls.2017.01620
Mascher M, Gundlach H, Himmelbach A, Beier S, Twardziok S O, Wicker T, Radchuk V, Dockter C, Hedley P E, Russell J, Bayer M, Ramsay L, Liu H, Haberer G, Zhang X-Q, Zhang Q, Barrero R A, Li L, Taudien S, Groth M, Felder M, Hastie A, Šimková H, Staňková H, Vrána J, Chan S, Muñoz-Amatriaín M, Ounit R, Wanamaker S, Bolser D, Colmsee C, Schmutzer T, Aliyeva-Schnorr L, Grasso S, Tanskanen J, Chailyan A, Sampath D, Heavens D, Clissold L, Cao S, Chapman B, Dai F, Han Y, Li H, Li X, Lin C, McCooke J K, Tan C, Wang P, Wang S, Yin S, Zhou G, Poland J A, Bellgard M I, Borisjuk L, Houben A, Doležel J, Ayling S, Lonardi S, Kersey P, Langridge P, Muehlbauer G J, Clark M D, Caccamo M, Schulman A H, Mayer K F X, Platzer M, Close T J, Scholz U, Hansson M, Zhang G, Braumann I, Spannagl M, Li C, Waugh R, Stein N:
A chromosome conformation capture ordered sequence of the barley genome. Nature 544 (2017) 427-433. https://dx.doi.org/10.1038/nature22043
Meier A, Worch S, Böer E, Hartmann A, Mascher M, Marzec M, Scholz U, Riechen J, Baronian K, Schauer F, Bode R, Kunze G:
Agdc1p – a gallic acid decarboxylase involved in the degradation of 1 tannic acid in the yeast Blastobotrys (Arxula) adeninivorans. Front. Microbiol. 8 (2017) 1777. https://dx.doi.org/10.3389/fmicb.2017.01777
Wendler N, Mascher M, Himmelbach A, Bini F, Kumlehn J, Stein N:
A high-density, sequence-enriched genetic map of Hordeum bulbosum and its collinearity to H. vulgare. Plant Genome 10 (2017) https://dx.doi.org/10.3835/plantgenome2017.06.0049
Wicker T, Schulman A H, Tanskanen J, Spannagl M, Twardziok S, Mascher M, Springer N M, Li Q, Waugh R, Li C, Zhang G, Stein N, Mayer K F X, Gundlach H:
The repetitive landscape of the 5100 Mbp barley genome. Mobile DNA 8 (2017) 22. https://dx.doi.org/10.1186/s13100-017-0102-3
Youssef H M, Mascher M, Ayoub M A, Stein N, Kilian B, Schnurbusch T:
Natural diversity of inflorescence architecture traces cryptic domestication genes in barley (Hordeum vulgare L.). Genet. Resour. Crop Evol. 64 (2017) 843-853. https://dx.doi.org/10.1007/s10722-017-0504-6
Beier S, Himmelbach A, Schmutzer T, Felder M, Taudien S, Mayer K F X, Platzer M, Stein N, Scholz U, Mascher M:
Multiplex sequencing of bacterial artificial chromosomes for assembling complex plant genomes. Plant Biotechnol. J. 14 (2016) 1511-1522. https://dx.doi.org/10.1111/pbi.12511
Jost M, Taketa S, Mascher M, Himmelbach A, Yuo T, Shahinnia F, Rutten T, Druka A, Schmutzer T, Steuernagel B, Beier S, Taudien S, Scholz U, Morgante M, Waugh R, Stein N:
A homolog of Blade-On-Petiole 1 and 2 (BOP1/2) controls internode length and homeotic changes of the barley inflorescence. Plant Physiol. 171 (2016) 1113-1127. https://dx.doi.org/10.1104/pp.16.00124
Livaja M, Unterseer S, Erath W, Lehermeier C, Wieseke R, Plieske J, Polley A, Luerssen H, Wieckhorst S, Mascher M, Hahn V, Ouzunova M, Schön C C, Ganal M W:
Diversity analysis and genomic prediction of Sclerotinia resistance in sunflower using a new 25 K SNP genotyping array. Theor. Appl. Genet. 129 (2016) 317-329. https://dx.doi.org/10.1007/s00122-015-2629-3
Mascher M, Schuenemann V J, Davidovich U, Marom N, Himmelbach A, Hübner S, Korol A, David M, Reiter E, Riehl S, Schreiber M, Vohr S H, Green R E, Dawson I K, Russell J, Kilian B, Muehlbauer G J, Waugh R, Fahima T, Krause J, Weiss E, Stein N:
Genomic analysis of 6,000-year-old cultivated grain illuminates the domestication history of barley. Nat. Genet. 48 (2016) 1089-1093. https://dx.doi.org/10.1038/ng.3611
Nagel M, Kodde J, Pistrick S, Mascher M, Börner A, Groot S P C:
Barley seed ageing: genetics behind the dry elevated pressure of oxygen ageing and moist controlled deterioration. Front. Plant Sci. 7 (2016) 388. https://dx.doi.org/10.3389/fpls.2016.00388
Rauter M, Kasprzak J, Becker K, Riechen J, Worch S, Hartmann A, Mascher M, Scholz U, Baronian K, Bode R, Schauer F, Matthias Vorbrodt H, Kunze G:
Aadh2p: an Arxula adeninivorans alcohol dehydrogenase involved in the first step of the 1-butanol degradation pathway. Microb. Cell Fact. 15 (2016) 175. https://dx.doi.org/10.1186/s12934-016-0573-9
Russell J, Mascher M, Dawson I K, Kyriakidis S, Calixto C, Freund F, Bayer M, Milne I, Marshall-Griffiths T, Heinen S, Hofstad A, Sharma R, Himmelbach A, Knauft M, van Zonneveld M, Brown J W, Schmid K, Kilian B, Muehlbauer G J, Stein N, Waugh R:
Exome sequencing of geographically diverse barley landraces and wild relatives gives insights into environmental adaptation. Nat. Genet. 48 (2016) 1024-1030. https://dx.doi.org/10.1038/ng.3612
Brown R H, Singh J, Singh S, Dahleen L S, Lemaux P G, Stein N, Mascher M, Bregitzer P:
Behavior of a modified Dissociation element in barley: a tool for genetic studies and for breeding transgenic barley. Mol. Breed. 35 (2015) 85. https://dx.doi.org/10.1007/s11032-015-0193-9
Chapman J A, Mascher M, Buluç A, Barry K, Georganas E, Session A, Strnadova V, Jenkins J, Seghal S, Oliker L, Schmutz J, Yelick K A, Scholz U, Waugh R, Poland J A, Muehlbauer G J, Stein N, Rokhsar D S:
A whole-genome shotgun approach for assembling and anchoring the hexaploid bread wheat genome. Genome Biol. 16 (2015) 26. https://dx.doi.org/10.1186/s13059-015-0582-8
Colmsee C, Beier S, Himmelbach A, Schmutzer T, Stein N, Scholz U, Mascher M:
BARLEX – the barley draft genome explorer. Mol. Plant 8 (2015) 964-966. https://dx.doi.org/10.1016/j.molp.2015.03.009
Lermontova I, Sandmann M, Mascher M, Schmit A C, Chabouté M E:
Centromeric chromatin and its dynamics in plants. Plant J. 83 (2015) 4-17. https://dx.doi.org/10.1111/tpj.12875
Mascher M:
Genetische Verankerung und Anordnung von Sequenzassemblies zur Unterstützung der genombasierten Pflanzenzüchtung. 17. Kurt von Rümker-Vorträge. Vortr. Pflanzenzücht. 85 (2015) 63-67.
Pourkheirandish M, Hensel G, Kilian B, Senthil N, Chen G, Sameri M, Azhaguvel P, Sakuma S, Dhanagond S, Sharma R, Mascher M, Himmelbach A, Gottwald S, Nair S K, Tagiri A, Yukuhiro F, Nagamura Y, Kanamori H, Matsumoto T, Willcox G, Middleton C P, Wicker T, Walther A, Waugh R, Fincher G B, Stein N, Kumlehn J, Sato K, Komatsuda T:
Evolution of the grain dispersal system in barley. Cell 162 (2015) 527-539. https://dx.doi.org/10.1016/j.cell.2015.07.002
Spannagl M, Alaux M, Lange M, Bolser D M, Bader K C, Letellier T, Kimmel E, Flores R, Pommier C, Kerhornou A, Walts B, Nussbaumer T, Grabmuller C, Chen J, Colmsee C, Beier S, Mascher M, Schmutzer T, Arend D, Thanki A, Ramirez-Gonzalez R, Ayling M, Ayling S, Caccamo M, Mayer K F X, Scholz U, Steinbach D, Quesneville H, Kersey P J:
transPLANT resources for Triticeae genomic data. Plant Genome 9 (2015) 1-13. https://dx.doi.org/10.3835/plantgenome2015.06.0038
Wendler N, Mascher M, Himmelbach A, Johnston P, Pickering R, Stein N:
Bulbosum to go: a toolbox to utilize Hordeum vulgare/ bulbosum introgressions for breeding and beyond. Mol. Plant 8 (2015) 1507-1519. https://dx.doi.org/10.1016/j.molp.2015.05.004
Zakhrabekova S, Dockter C, Ahmann K, Braumann I, Gough S P, Wendt T, Lundqvist U, Mascher M, Stein N, Hansson M:
Genetic linkage facilitates cloning of Ert-m regulating plant architecture in barley and identified a strong candidate of Ant1 involved in anthocyanin biosynthesis. Plant Mol. Biol. 88 (2015) 609-626. https://dx.doi.org/10.1007/s11103-015-0350-x
Datenpublikationen
Avni R, Mascher M:
PepsiCo OT3098 hexaploid oat version 2 genome assembly. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB76239 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB76239
Guo Y:
A haplotype-based evolutionary history of barley domestication. European Variation Archive (EVA) at EMBL-EBI under accession number PRJEB79752 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB79752
Guo Y:
Ancestral haplotype groups in a diversity panel of wild and domesticated barley (Hordeum vulgare). e!DAL - Plant Genomics and Phenomics Research Data Repository (PGP), IPK Gatersleben (2025) https://dx.doi.org/10.5447/IPK/2025/7
Guo Y, Mascher M:
Wild barley diversity panel. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB65046 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB65046
Haupt M:
AGENT: Atlas of barley. European Variation Archive (EVA) at EMBL-EBI under accession number PRJEB94862 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB94862
Mascher M:
Global genomic diversity of hexaploid oat. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB63645 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB63645
Mascher M:
Exome capture sequencing data for barley (Hordeum vulgare) gravitropism mutant TM194. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB44451 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB44451
Mascher M:
Chromosome-scale assembly of Avena byzantina accession PI 258586. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB45067 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB45067
Mascher M:
Avena sativa cv. Leggett assembly. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB62638 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB62638
Mascher M:
Avena sativa cv. Morgan assembly. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB62639 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB62639
Mascher M:
Avena sativa cv. Williams assembly. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB62640 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB62640
Mascher M, Feng J-W:
Pangenome graphs and variant matrix derived from haplotype-resolved genome sequence assemblies of Hordeum bulbosum. e!DAL - Plant Genomics and Phenomics Research Data Repository (PGP), IPK Gatersleben (2025) https://dx.doi.org/10.5447/IPK/2025/4
PanOat, Mascher M:
Avena sativa cv. Delfin genome assembly. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB56707 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB56707
PanOat, Mascher M:
Avena sativa cv. Victoria genome assembly. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB56706 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB56706
PanOat, Mascher M:
Avena sativa OT380 genome assembly. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB56705 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB56705
PanOat, Mascher M:
Avena sativa cv. Amagalon genome assembly. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB56704 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB56704
PanOat, Mascher M:
Avena sativa FM13 genome assembly. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB56703 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB56703
PanOat, Mascher M:
Avena sativa cv. Rhapsody genome assembly. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB56702 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB56702
PanOat, Mascher M:
Avena sativa cv. Lion genome assembly. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB56701 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB56701
PanOat, Mascher M:
Avena sativa cv. Bingo genome assembly. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB56700 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB56700
PanOat, Mascher M:
Avena abyssinica BYU960 genome assembly. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB56724 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB56724
PanOat, Mascher M:
Avena barbata accession PI388828 genome assembly. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB56723 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB56723
PanOat, Mascher M:
Avena sterilis TN4 genome assembly. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB56722 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB56722
PanOat, Mascher M:
Avena sativa cv. HiFi genome assembly. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB56721 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB56721
PanOat, Mascher M:
Avena sativa cv. Gehl genome assembly. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB56720 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB56720
PanOat, Mascher M:
Avena sativa cv. GS7 genome assembly. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB56719 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB56719
PanOat, Mascher M:
Avena sativa cv. Clintland60 genome assembly. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB56718 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB56718
PanOat, Mascher M:
Avena sativa cv. Aslak genome assembly. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB56717 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB56717
PanOat, Mascher M:
Avena sterilis TN1 genome assembly. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB56716 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB56716
PanOat, Mascher M:
Avena sativa accession PI182478 genome assembly. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB56715 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB56715
PanOat, Mascher M:
Avena sativa cv. Park genome assembly. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB56714 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB56714
PanOat, Mascher M:
Avena fatua CN25955 genome assembly. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB56713 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB56713
PanOat, Mascher M:
Avena sativa cv. Bannister genome assembly. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB56712 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB56712
PanOat, Mascher M:
Avena sativa GMI423 genome assembly. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB56711 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB56711
PanOat, Mascher M:
Avena sativa cv. Hatives des Alpes genome assembly. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB56710 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB56710
PanOat, Mascher M:
Avena sativa cv. Bilby genome assembly. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB56709 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB56709
PanOat, Mascher M:
Avena sativa cv. Nicolas genome assembly. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB56708 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB56708
PanOat, Mascher M:
PanOat project: Raw data. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB56828 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB56828
PanOat, Mascher M:
Transcriptome data of diverse oats (Avena spp.). European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB57570 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB57570
PanOat, Mascher M:
PanOat:WGS Resequencing. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB62778 (2025) https://www.ebi.ac.uk/ena/browser/view/PRJEB62778
Avni R, Mascher M:
Aegilops longissima - long read genome assembly. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB58051 (2024) https://www.ebi.ac.uk/ena/browser/view/PRJEB58051
Avni R, Mascher M:
A single NLR gene confers resistance to both leaf and stripe rust in wheat (RNA-Seq). European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB64238 (2024) https://www.ebi.ac.uk/ena/browser/view/PRJEB64238
Avni R, Mascher M:
A single NLR gene confers resistance to both leaf and stripe rust in wheat (RenSeq). European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB64239 (2024) https://www.ebi.ac.uk/ena/browser/view/PRJEB64239
Chen Y, Kölliker R, Mascher M, Copetti D, Himmelbach A, Stein N, Studer B:
Supporting data for An improved chromosome-level genome assembly of perennial ryegrass (Lolium perenne L.)"." GigaScience Database (2024) https://dx.doi.org/10.5524/102500
Cho W, Mascher M:
Genome assemblies of a diploid chamomile (Matricaria recutita) genotype. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB64706 (2024) https://www.ebi.ac.uk/ena/browser/view/PRJEB64706
Cho W, Otto L G, Mascher M:
An annotated haplotype-resolved genome sequence assembly of diploid German chamomile, Matricaria chamomilla. NCBI GenBank under accession number PRJNA1147238 (2024) https://www.ncbi.nlm.nih.gov/bioproject/1147238
Guo Y, Mascher M:
Single-nucleotide polymorphism matrices for 281 wild barleys (Hordeum vulgare) subsp. spontaneum from the Wild Barley Diversity Collection (WBDC). European Variation Archive (EVA) at EMBL-EBI under accession number PRJEB80165 (2024) https://www.ebi.ac.uk/ena/browser/view/PRJEB80165
Haupt M:
Major chromosome rearrangements in intergeneric wheat x rye hybrids in compatible and incompatible crosses detected by GBS read coverage analysis. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB64645 (2024) https://www.ebi.ac.uk/ena/data/view/PRJEB64645
Haupt M, Tikhenko N, Mascher M:
Major chromosome rearrangements in intergeneric wheat x rye hybrids in compatible and incompatible crosses detected by GBS read coverage analysis. e!DAL - Plant Genomics and Phenomics Research Data Repository (PGP), IPK Gatersleben (2024) https://dx.doi.org/10.5447/IPK/2024/4
Jayakodi M, Mascher M:
SNP matrix of barley core set from german ex-situ genbank IPK selected for BPGv2 project. European Variation Archive (EVA) at EMBL-EBI under accession number PRJEB70778 (2024) https://www.ebi.ac.uk/ena/browser/view/PRJEB70778
Mascher M:
WBDC-WGS. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB56087 (2024) https://www.ebi.ac.uk/ena/browser/view/PRJEB56087
Mascher M:
A pangenome of the barley wild relative Hordeum bulbosum. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB65276 (2024) https://www.ebi.ac.uk/ena/browser/view/PRJEB65276
Mascher M:
Single pollen nuclei sequencing data of Hordeum bulbosum clone FB19-011-3. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB65918 (2024) https://www.ebi.ac.uk/ena/browser/view/PRJEB65918
Stein N, Mascher M:
BPGv2: HiFi and Hi-C raw data. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB57567 (2024) https://www.ebi.ac.uk/ena/data/view/PRJEB57567
Câmara* A, Schubert* V:
Helical organization of barley metaphase chromatids. e!DAL - Plant Genomics and Phenomics Research Data Repository (PGP), IPK Gatersleben (2023) https://dx.doi.org/10.5447/IPK/2023/2
El Hanafi* S, Jiang* Y, Reif* J, Kehel Z, Schulthess Börgel A W, Zhao Y, Mascher M, Haupt M, Himmelbach A, Stein N, Amri A:
Genome-wide prediction of thousand kernel weight for 234 winter barley accessions from the ICARDA genebank using 1,910 IPK genebank accessions as training set. e!DAL - Plant Genomics and Phenomics Research Data Repository (PGP), IPK Gatersleben (2023) https://dx.doi.org/10.5447/IPK/2023/8
Kale S M, Mascher M:
Raw sequence data of 47 diverse barley (Hordeum vulgare L.) genotypes. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB45512 (2023) https://www.ebi.ac.uk/ena/data/view/PRJEB45512
Avni R, Mascher M:
Chromosome-scale sequence assembly of the genome of bread wheat (Triticum aestivum) cultivar Attraktion. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB48529 (2022) https://www.ebi.ac.uk/ena/data/view/PRJEB48529
Avni R, Mascher M:
10X Genomics Chromium reads for Aegilops speltoides var. speltoides accession Tivon. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB48314 (2022) https://www.ebi.ac.uk/ena/data/view/PRJEB48314
Beier S, Mascher M:
BRIDGE-MorexV3: SNP matrix of barley collection from german ex-situ genbank IPK. European Variation Archive (EVA) at EMBL-EBI under accession number PRJEB51851 (2022) https://www.ebi.ac.uk/eva/?eva-study=PRJEB51851
Guo Y, Mascher M:
Variant matrices for six-rowed wild-growing barleys (Hordeum agriocrithon). e!DAL - Plant Genomics and Phenomics Research Data Repository (PGP), IPK Gatersleben (2022) https://dx.doi.org/10.5447/IPK/2022/10
Guo Y, Mascher M:
Wild Barley H. agriocrition GBS, WGS3x, WGS10x. European Variation Archive (EVA) at EMBL-EBI under accession number PRJEB53985 (2022) https://www.ebi.ac.uk/eva/?eva-study=PRJEB53985
Huang Y, Schnurbusch T, Mascher M:
Diversity of panel of wild and domesticated barley (Hordeum vulgare) and crop-wild hybrids. European Variation Archive (EVA) at EMBL-EBI under accession number PRJEB62782 (2022) https://www.ebi.ac.uk/eva/?eva-study=PRJEB62782
Kale S M:
Genebank 2.0 - Resistance gene capture in winter wheat diversity panel. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB48219 (2022) https://www.ebi.ac.uk/ena/data/view/PRJEB48219
Kale S M, Mascher M:
Genebank 2.0: Resistance gene capture in winter wheat diversity panels. European Variation Archive (EVA) at EMBL-EBI under accession number PRJEB52597 (2022) https://www.ebi.ac.uk/eva/?eva-study=PRJEB52597
Mascher M:
Filtration script for genetic variant matrices in Variant Call Format (VCF). e!DAL - Plant Genomics and Phenomics Research Data Repository (PGP), IPK Gatersleben (2022) https://dx.doi.org/10.5447/IPK/2022/15
Mascher M:
Genotyping-by-sequencing data of the Wild Barley Diversity Collection. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB51697 (2022) https://www.ebi.ac.uk/ena/data/view/PRJEB51697
Mascher M:
Six-rowed wild-growing barley GBS. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB51488 (2022) https://www.ebi.ac.uk/ena/data/view/PRJEB51488
Mascher M:
Six-rowed wild-growing barley WGS. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB51487 (2022) https://www.ebi.ac.uk/ena/data/view/PRJEB51487
Mascher M:
Genotyping-by-sequencing data of a rye (Secale spp.) diversity panel. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB50548 (2022) https://www.ebi.ac.uk/ena/data/view/PRJEB50548
Mascher M:
Recombination landscape divergence between populations is marked by larger low-recombining regions in domesticated rye. European Variation Archive (EVA) at EMBL-EBI under accession number PRJEB51528 (2022) https://www.ebi.ac.uk/eva/?eva-study=PRJEB51528
Mascher M:
Whole-genome re-sequencing of barley cultivar Chikurin Ibaraki 1. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB50079 (2022) https://www.ebi.ac.uk/ena/data/view/PRJEB50079
Mascher M, Marone M:
TRITEX pipeline source code and documentation. e!DAL - Plant Genomics and Phenomics Research Data Repository (PGP), IPK Gatersleben (2022) https://dx.doi.org/10.5447/IPK/2022/28
Püpke Marone M, Mascher M:
Example files generated in the TRITEX long-read assembly pipeline. e!DAL - Plant Genomics and Phenomics Research Data Repository (PGP), IPK Gatersleben (2022) https://dx.doi.org/10.5447/IPK/2022/20
Schulthess A, Mascher M:
Single-nucleotide polymorphism matrices for a large diversity panel of wheat (Triticum aestivum L.). European Variation Archive (EVA) at EMBL-EBI under accession number PRJEB52759 (2022) https://www.ebi.ac.uk/eva/?eva-study=PRJEB52759
Schulthess A W, Kale S M, Liu F, Zhao Y, Philipp N, Rembe M, Jiang Y, Beukert U, Serfling A, Himmelbach A, Fuchs J, Oppermann M, Weise S, Boeven P H G, Schacht J, Longin C F H, Kollers S, Pfeiffer N, Korzun V, Lange M, Scholz U, Stein N, Mascher M, Reif J C:
Genome-wide association mapping for yellow rust resistance in a population of 454 whole-genome sequenced diverse wheat genotypes. e!DAL - Plant Genomics and Phenomics Research Data Repository (PGP), IPK Gatersleben (2022) https://dx.doi.org/10.5447/ipk/2022/5
Schulthess A W, Kale S M, Liu F, Zhao Y, Philipp N, Rembe M, Jiang Y, Beukert U, Serfling A, Himmelbach A, Fuchs J, Oppermann M, Weise S, Boeven P H G, Schacht J, Longin C F H, Kollers S, Pfeiffer N, Korzun V, Lange M, Scholz U, Stein N, Mascher M, Reif J C:
Genomic prediction of yield breeding values for 10,353 winter wheat genebank samples. e!DAL - Plant Genomics and Phenomics Research Data Repository (PGP), IPK Gatersleben (2022) https://dx.doi.org/10.5447/ipk/2022/6
Schulthess A W, Kale S M, Zhao Y, Gogna A, Rembe M, Philipp N, Liu F, Beukert U, Serfling A, Himmelbach A, Oppermann M, Weise S, Boeven P H G, Schacht J, Longin C F H, Kollers S, Pfeiffer N, Korzun V, Fiebig A, Schüler D, Lange M, Scholz U, Stein N, Mascher M, Reif J C:
Genomic-phenotypic data interoperability between 8,838 genotyped wheat samples, grain yield breeding value estimates and yellow rust infection scores from multiple-environment field trials. e!DAL - Plant Genomics and Phenomics Research Data Repository (PGP), IPK Gatersleben (2022) https://dx.doi.org/10.5447/IPK/2022/21
Schulthess A W, Kale S M, Zhao Y, Gogna A, Rembe M, Philipp N, Liu F, Beukert U, Serfling A, Himmelbach A, Oppermann M, Weise S, Boeven P H G, Schacht J, Longin C F H, Kollers S, Pfeiffer N, Korzun V, Fiebig A, Schüler D, Lange M, Scholz U, Stein N, Mascher M, Reif J C:
Evaluating the yellow rust resistance of 7,682 winter wheat IPK genebank accessions and 80 modern European cultivars based on natural infections in multiple-environments. e!DAL - Plant Genomics and Phenomics Research Data Repository (PGP), IPK Gatersleben (2022) https://dx.doi.org/10.5447/IPK/2022/22
Schulthess A W, Kale S M, Zhao Y, Gogna A, Rembe M, Philipp N, Liu F, Beukert U, Serfling A, Himmelbach A, Oppermann M, Weise S, Boeven P H G, Schacht J, Longin C F H, Kollers S, Pfeiffer N, Korzun V, Fiebig A, Schüler D, Lange M, Scholz U, Stein N, Mascher M, Reif J C:
Evaluating the yellow rust resistance of 600 winter wheat IPK genebank accession samples and 199 modern European cultivars based on natural and artificial inoculations in multiple-environments. e!DAL - Plant Genomics and Phenomics Research Data Repository (PGP), IPK Gatersleben (2022) https://dx.doi.org/10.5447/IPK/2022/23
Schulthess A W, Kale S M, Zhao Y, Gogna A, Rembe M, Philipp N, Liu F, Beukert U, Serfling A, Himmelbach A, Oppermann M, Weise S, Boeven P H G, Schacht J, Longin C F H, Kollers S, Pfeiffer N, Korzun V, Fiebig A, Schüler D, Lange M, Scholz U, Stein N, Mascher M, Reif J C:
Multiple-environment yield evaluation of 173 advanced wheat pre-breeding lines from crosses involving IPK genebank accessions with high yield breeding values. e!DAL - Plant Genomics and Phenomics Research Data Repository (PGP), IPK Gatersleben (2022) https://dx.doi.org/10.5447/IPK/2022/24
Schulthess A W, Kale S M, Zhao Y, Gogna A, Rembe M, Philipp N, Liu F, Beukert U, Serfling A, Himmelbach A, Oppermann M, Weise S, Boeven P H G, Schacht J, Longin C F H, Kollers S, Pfeiffer N, Korzun V, Fiebig A, Schüler D, Lange M, Scholz U, Stein N, Mascher M, Reif J C:
Passport information for 8,838 genotyped wheat samples. e!DAL - Plant Genomics and Phenomics Research Data Repository (PGP), IPK Gatersleben (2022) https://dx.doi.org/10.5447/ipk/2022/31
Schulthess A W, Mascher M:
Genebank 2.0 - GENDIV. European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB48988 (2022) https://www.ebi.ac.uk/ena/data/view/PRJEB48988
Zhao Y, Schulthess A W, Kale S M, Gogna A, Rembe M, Philipp N, Liu F, Beukert U, Serfling A, Himmelbach A, Oppermann M, Weise S, Boeven P H G, Schacht J, Longin C F H, Kollers S, Pfeiffer N, Korzun V, Fiebig A, Schüler D, Lange M, Scholz U, Stein N, Mascher M, Reif J C:
Estimating the breeding values of 707 winter wheat IPK genebank accessions using yields of 'Elite×PGR' F1 hybrids tested across multiple-environment experiments. e!DAL - Plant Genomics and Phenomics Research Data Repository (PGP), IPK Gatersleben (2022) https://dx.doi.org/10.5447/IPK/2022/25
scroll top